Search Results for author: Gabriele Corso
Found 16 papers, 14 papers with code
Deep Confident Steps to New Pockets: Strategies for Docking Generalization
2 code implementations • 28 Feb 2024 • Gabriele Corso, Arthur Deng, Benjamin Fry, Nicholas Polizzi, Regina Barzilay, Tommi Jaakkola
Accurate blind docking has the potential to lead to new biological breakthroughs, but for this promise to be realized, docking methods must generalize well across the proteome.
Dirichlet Flow Matching with Applications to DNA Sequence Design
1 code implementation • 8 Feb 2024 • Hannes Stark, Bowen Jing, Chenyu Wang, Gabriele Corso, Bonnie Berger, Regina Barzilay, Tommi Jaakkola
Further, we provide distilled Dirichlet flow matching, which enables one-step sequence generation with minimal performance hits, resulting in $O(L)$ speedups compared to autoregressive models.
Particle Guidance: non-I.I.D. Diverse Sampling with Diffusion Models
1 code implementation • 19 Oct 2023 • Gabriele Corso, Yilun Xu, Valentin De Bortoli, Regina Barzilay, Tommi Jaakkola
In light of the widespread success of generative models, a significant amount of research has gone into speeding up their sampling time.
DiffDock-PP: Rigid Protein-Protein Docking with Diffusion Models
1 code implementation • 8 Apr 2023 • Mohamed Amine Ketata, Cedrik Laue, Ruslan Mammadov, Hannes Stärk, Menghua Wu, Gabriele Corso, Céline Marquet, Regina Barzilay, Tommi S. Jaakkola
Understanding how proteins structurally interact is crucial to modern biology, with applications in drug discovery and protein design.
EigenFold: Generative Protein Structure Prediction with Diffusion Models
1 code implementation • 5 Apr 2023 • Bowen Jing, Ezra Erives, Peter Pao-Huang, Gabriele Corso, Bonnie Berger, Tommi Jaakkola
Protein structure prediction has reached revolutionary levels of accuracy on single structures, yet distributional modeling paradigms are needed to capture the conformational ensembles and flexibility that underlie biological function.
Modeling Molecular Structures with Intrinsic Diffusion Models
1 code implementation • 23 Feb 2023 • Gabriele Corso
Since its foundations, more than one hundred years ago, the field of structural biology has strived to understand and analyze the properties of molecules and their interactions by studying the structure that they take in 3D space.
Learning Graph Search Heuristics
no code implementations • Learning on Graphs 2022 • Michal Pándy, Weikang Qiu, Gabriele Corso, Petar Veličković, Rex Ying, Jure Leskovec, Pietro Liò
At training time, we aggregate datasets of search trajectories and ground-truth shortest path distances, which we use to train a specialized graph neural network-based heuristic function using backpropagation through steps of the pathfinding process.
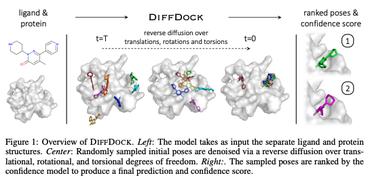
DiffDock: Diffusion Steps, Twists, and Turns for Molecular Docking
2 code implementations • 4 Oct 2022 • Gabriele Corso, Hannes Stärk, Bowen Jing, Regina Barzilay, Tommi Jaakkola
We instead frame molecular docking as a generative modeling problem and develop DiffDock, a diffusion generative model over the non-Euclidean manifold of ligand poses.
 Ranked #1 on
Blind Docking
on PDBbind
Ranked #1 on
Blind Docking
on PDBbind
Torsional Diffusion for Molecular Conformer Generation
1 code implementation • 1 Jun 2022 • Bowen Jing, Gabriele Corso, Jeffrey Chang, Regina Barzilay, Tommi Jaakkola
Molecular conformer generation is a fundamental task in computational chemistry.
Subspace Diffusion Generative Models
1 code implementation • 3 May 2022 • Bowen Jing, Gabriele Corso, Renato Berlinghieri, Tommi Jaakkola
Score-based models generate samples by mapping noise to data (and vice versa) via a high-dimensional diffusion process.
 Ranked #19 on
Image Generation
on CIFAR-10
Ranked #19 on
Image Generation
on CIFAR-10
Graph Anisotropic Diffusion
1 code implementation • 30 Apr 2022 • Ahmed A. A. Elhag, Gabriele Corso, Hannes Stärk, Michael M. Bronstein
Traditional Graph Neural Networks (GNNs) rely on message passing, which amounts to permutation-invariant local aggregation of neighbour features.
3D Infomax improves GNNs for Molecular Property Prediction
1 code implementation • NeurIPS Workshop AI4Scien 2021 • Hannes Stärk, Dominique Beaini, Gabriele Corso, Prudencio Tossou, Christian Dallago, Stephan Günnemann, Pietro Liò
Molecular property prediction is one of the fastest-growing applications of deep learning with critical real-world impacts.
3D Pre-training improves GNNs for Molecular Property Prediction
no code implementations • 29 Sep 2021 • Hannes Stärk, Dominique Beaini, Gabriele Corso, Prudencio Tossou, Christian Dallago, Stephan Günnemann, Pietro Lio
Molecular property prediction is one of the fastest-growing applications of deep learning with critical real-world impacts.
Neural Distance Embeddings for Biological Sequences
1 code implementation • NeurIPS 2021 • Gabriele Corso, Rex Ying, Michal Pándy, Petar Veličković, Jure Leskovec, Pietro Liò
The development of data-dependent heuristics and representations for biological sequences that reflect their evolutionary distance is critical for large-scale biological research.
Directional Graph Networks
2 code implementations • 6 Oct 2020 • Dominique Beaini, Saro Passaro, Vincent Létourneau, William L. Hamilton, Gabriele Corso, Pietro Liò
Then, we propose the use of the Laplacian eigenvectors as such vector field.
 Ranked #2 on
Node Classification
on PATTERN 100k
Ranked #2 on
Node Classification
on PATTERN 100k
Principal Neighbourhood Aggregation for Graph Nets
8 code implementations • NeurIPS 2020 • Gabriele Corso, Luca Cavalleri, Dominique Beaini, Pietro Liò, Petar Veličković
Graph Neural Networks (GNNs) have been shown to be effective models for different predictive tasks on graph-structured data.
 Ranked #4 on
Node Classification
on PATTERN 100k
Ranked #4 on
Node Classification
on PATTERN 100k