Bayesian Hybrid Matrix Factorisation for Data Integration
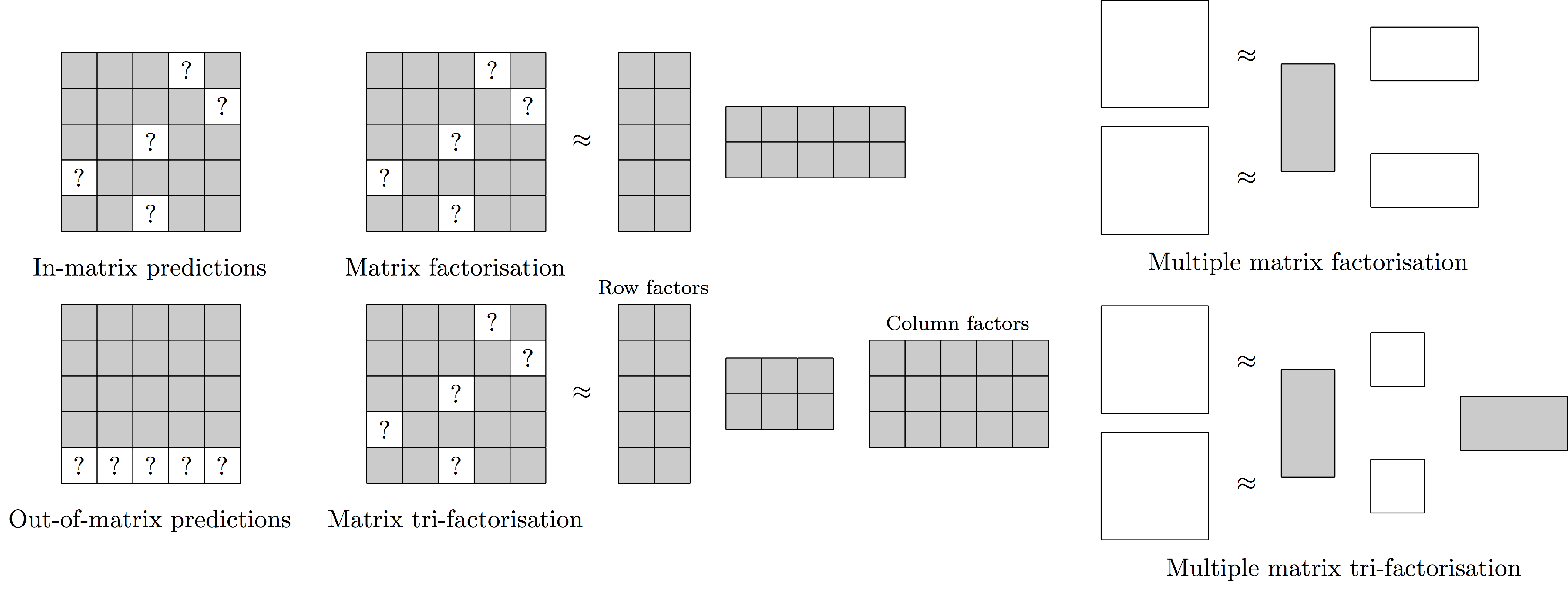
We introduce a novel Bayesian hybrid matrix factorisation model (HMF) for data integration, based on combining multiple matrix factorisation methods, that can be used for in- and out-of-matrix prediction of missing values. The model is very general and can be used to integrate many datasets across different entity types, including repeated experiments, similarity matrices, and very sparse datasets. We apply our method on two biological applications, and extensively compare it to state-of-the-art machine learning and matrix factorisation models. For in-matrix predictions on drug sensitivity datasets we obtain consistently better performances than existing methods. This is especially the case when we increase the sparsity of the datasets. Furthermore, we perform out-of-matrix predictions on methylation and gene expression datasets, and obtain the best results on two of the three datasets, especially when the predictivity of datasets is high.
PDF Abstract