BioELECTRA:Pretrained Biomedical text Encoder using Discriminators
Recent advancements in pretraining strategies in NLP have shown a significant improvement in the performance of models on various text mining tasks. We apply ‘replaced token detection’ pretraining technique proposed by ELECTRA and pretrain a biomedical language model from scratch using biomedical text and vocabulary. We introduce BioELECTRA, a biomedical domain-specific language encoder model that adapts ELECTRA for the Biomedical domain. WE evaluate our model on the BLURB and BLUE biomedical NLP benchmarks. BioELECTRA outperforms the previous models and achieves state of the art (SOTA) on all the 13 datasets in BLURB benchmark and on all the 4 Clinical datasets from BLUE Benchmark across 7 different NLP tasks. BioELECTRA pretrained on PubMed and PMC full text articles performs very well on Clinical datasets as well. BioELECTRA achieves new SOTA 86.34%(1.39% accuracy improvement) on MedNLI and 64% (2.98% accuracy improvement) on PubMedQA dataset.
PDF AbstractCode
Results from the Paper
| Task | Dataset | Model | Metric Name | Metric Value | Global Rank | Benchmark |
|---|---|---|---|---|---|---|
| Natural Language Inference | MedNLI | BioELECTRA-Base | Accuracy | 86.34 | # 2 | |
| Question Answering | PubMedQA | BioELECTRA uncased | Accuracy | 64.2 | # 21 | |
| Medical Named Entity Recognition | ShARe/CLEF eHealth corpus | BioELECTRA | F1 | 0.8371 | # 1 |







 SQuAD
SQuAD
 BC5CDR
BC5CDR
 BioASQ
BioASQ
 PubMedQA
PubMedQA
 DDI
DDI
 BIOSSES
BIOSSES
 BLURB
BLURB
 HOC
HOC
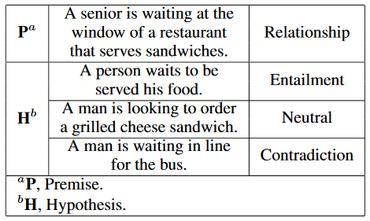
 MedNLI
MedNLI
