Conditional molecular design with deep generative models
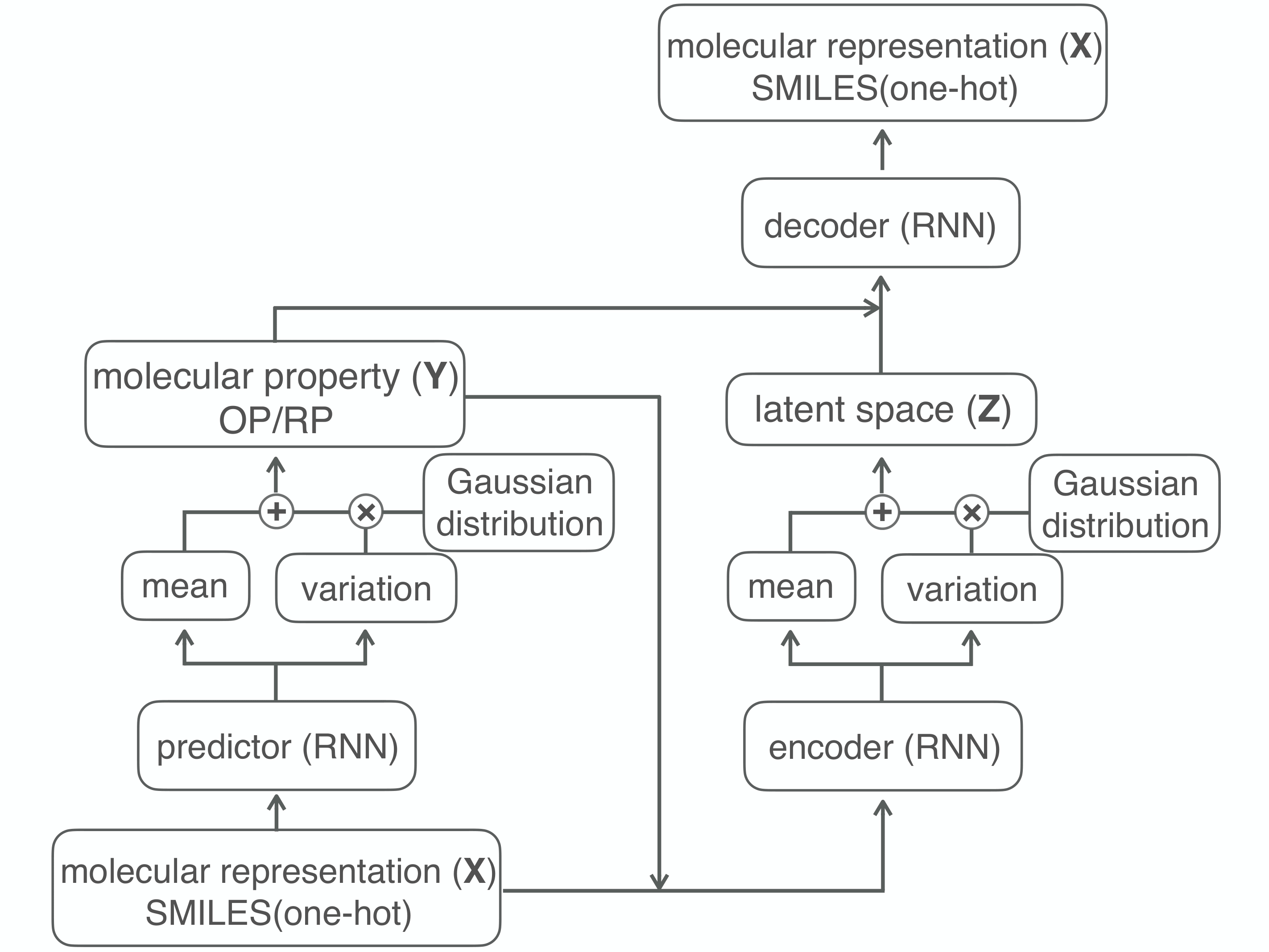
Although machine learning has been successfully used to propose novel molecules that satisfy desired properties, it is still challenging to explore a large chemical space efficiently. In this paper, we present a conditional molecular design method that facilitates generating new molecules with desired properties. The proposed model, which simultaneously performs both property prediction and molecule generation, is built as a semi-supervised variational autoencoder trained on a set of existing molecules with only a partial annotation. We generate new molecules with desired properties by sampling from the generative distribution estimated by the model. We demonstrate the effectiveness of the proposed model by evaluating it on drug-like molecules. The model improves the performance of property prediction by exploiting unlabeled molecules, and efficiently generates novel molecules fulfilling various target conditions.
PDF Abstract