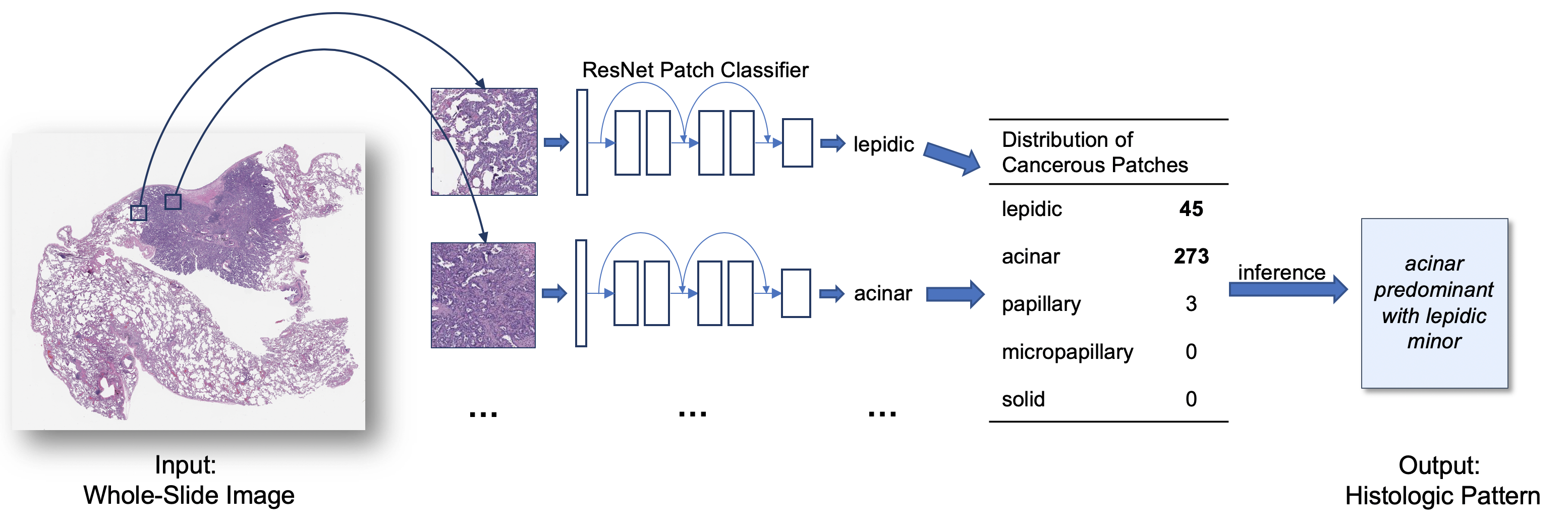
Attention-Based Deep Neural Networks for Detection of Cancerous and Precancerous Esophagus Tissue on Histopathological Slides
Deep learning-based methods, such as the sliding window approach for cropped-image classification and heuristic aggregation for whole-slide inference, for analyzing histological patterns in high-resolution microscopy images have shown promising results. These approaches, however, require a laborious annotation process and are fragmented. This diagnostic study collected deidentified high-resolution histological images (N = 379) for training a new model composed of a convolutional neural network and a grid-based attention network, trainable without region-of-interest annotations. Histological images of patients who underwent endoscopic esophagus and gastroesophageal junction mucosal biopsy between January 1, 2016, and December 31, 2018, at Dartmouth-Hitchcock Medical Center (Lebanon, New Hampshire) were collected. The method achieved a mean accuracy of 0.83 in classifying 123 test images. These results were comparable with or better than the performance from the current state-of-the-art sliding window approach, which was trained with regions of interest. Results of this study suggest that the proposed attention-based deep neural network framework for Barrett esophagus and esophageal adenocarcinoma detection is important because it is based solely on tissue-level annotations, unlike existing methods that are based on regions of interest. This new model is expected to open avenues for applying deep learning to digital pathology.
PDF AbstractCode
Datasets
Results from the Paper
| Task | Dataset | Model | Metric Name | Metric Value | Global Rank | Benchmark |
|---|---|---|---|---|---|---|
| Medical Object Detection | Barrett’s Esophagus | Attention-based model | Mean Accuracy | 81% | # 1 |