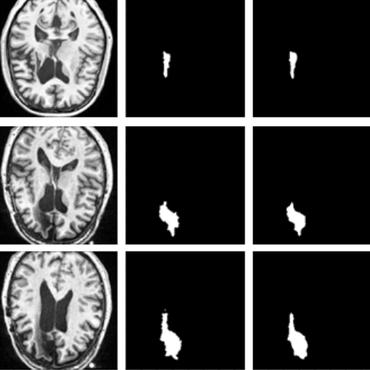
Learning joint segmentation of tissues and brain lesions from task-specific hetero-modal domain-shifted datasets
Brain tissue segmentation from multimodal MRI is a key building block of many neuroimaging analysis pipelines. Established tissue segmentation approaches have, however, not been developed to cope with large anatomical changes resulting from pathology, such as white matter lesions or tumours, and often fail in these cases. In the meantime, with the advent of deep neural networks (DNNs), segmentation of brain lesions has matured significantly. However, few existing approaches allow for the joint segmentation of normal tissue and brain lesions. Developing a DNN for such a joint task is currently hampered by the fact that annotated datasets typically address only one specific task and rely on task-specific imaging protocols including a task-specific set of imaging modalities. In this work, we propose a novel approach to build a joint tissue and lesion segmentation model from aggregated task-specific hetero-modal domain-shifted and partially-annotated datasets. Starting from a variational formulation of the joint problem, we show how the expected risk can be decomposed and optimised empirically. We exploit an upper bound of the risk to deal with heterogeneous imaging modalities across datasets. To deal with potential domain shift, we integrated and tested three conventional techniques based on data augmentation, adversarial learning and pseudo-healthy generation. For each individual task, our joint approach reaches comparable performance to task-specific and fully-supervised models. The proposed framework is assessed on two different types of brain lesions: White matter lesions and gliomas. In the latter case, lacking a joint ground-truth for quantitative assessment purposes, we propose and use a novel clinically-relevant qualitative assessment methodology.
PDF Abstract