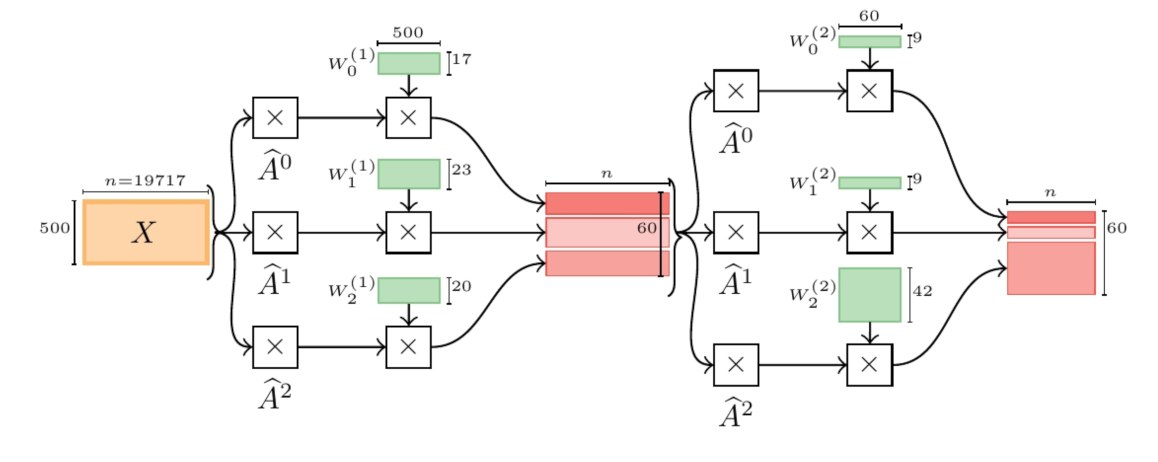
MixHop: Higher-Order Graph Convolutional Architectures via Sparsified Neighborhood Mixing
Existing popular methods for semi-supervised learning with Graph Neural Networks (such as the Graph Convolutional Network) provably cannot learn a general class of neighborhood mixing relationships. To address this weakness, we propose a new model, MixHop, that can learn these relationships, including difference operators, by repeatedly mixing feature representations of neighbors at various distances. Mixhop requires no additional memory or computational complexity, and outperforms on challenging baselines. In addition, we propose sparsity regularization that allows us to visualize how the network prioritizes neighborhood information across different graph datasets. Our analysis of the learned architectures reveals that neighborhood mixing varies per datasets.
PDF AbstractCode
Datasets
 Cora
Citeseer
Cora
Citeseer
 Wiki Squirrel
WebKB
Penn94
Cornell
genius
twitch-gamers
Deezer-Europe
Chameleon (48%/32%/20% fixed splits)
Squirrel (60%/20%/20% random splits)
Squirrel (48%/32%/20% fixed splits)
Wisconsin(60%/20%/20% random splits)
Cornell (48%/32%/20% fixed splits)
PubMed (60%/20%/20% random splits)
Cornell (60%/20%/20% random splits)
Texas(60%/20%/20% random splits)
Film (60%/20%/20% random splits)
Citeseer (48%/32%/20% fixed splits)
PubMed (48%/32%/20% fixed splits)
Wisconsin (48%/32%/20% fixed splits)
Chameleon(60%/20%/20% random splits)
Cora (48%/32%/20% fixed splits)
Texas (48%/32%/20% fixed splits)
Film(48%/32%/20% fixed splits)
Wiki Squirrel
WebKB
Penn94
Cornell
genius
twitch-gamers
Deezer-Europe
Chameleon (48%/32%/20% fixed splits)
Squirrel (60%/20%/20% random splits)
Squirrel (48%/32%/20% fixed splits)
Wisconsin(60%/20%/20% random splits)
Cornell (48%/32%/20% fixed splits)
PubMed (60%/20%/20% random splits)
Cornell (60%/20%/20% random splits)
Texas(60%/20%/20% random splits)
Film (60%/20%/20% random splits)
Citeseer (48%/32%/20% fixed splits)
PubMed (48%/32%/20% fixed splits)
Wisconsin (48%/32%/20% fixed splits)
Chameleon(60%/20%/20% random splits)
Cora (48%/32%/20% fixed splits)
Texas (48%/32%/20% fixed splits)
Film(48%/32%/20% fixed splits)
Results from the Paper
| Task | Dataset | Model | Metric Name | Metric Value | Global Rank | Benchmark |
|---|---|---|---|---|---|---|
| Node Classification | Actor | MixHop | Accuracy | 32.22 ± 2.34 | # 42 | |
| Node Classification | Chameleon | MixHop | Accuracy | 60.50 ± 2.53 | # 46 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Chameleon (48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 60.50 ± 2.53 | # 23 | |
| Node Classification | Chameleon (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 36.28 ± 10.22 | # 36 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Chameleon(60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 36.28 ± 10.22 | # 32 | |
| Node Classification | Citeseer | MixHop | Accuracy | 71.4% | # 56 | |
| Training Split | 20 per node | # 1 | ||||
| Validation | YES | # 1 | ||||
| Node Classification | Citeseer (48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 76.26 ± 1.33 | # 20 | |
| Node Classification | CiteSeer (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 49.52 ± 13.35 | # 32 | |
| Node Classification | Cora | MixHop | Accuracy | 81.9% | # 58 | |
| Training Split | 20 per node | # 1 | ||||
| Validation | YES | # 1 | ||||
| Node Classification | Cora (48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 87.61 ± 0.85 | # 17 | |
| Node Classification | Cora (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 65.65 ± 11.31 | # 32 | |
| Node Classification | Cornell | MixHop | Accuracy | 73.51 ± 6.34 | # 43 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Cornell (48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 73.51 ± 6.34 | # 24 | |
| Node Classification | Cornell (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 60.33 ± 28.53 | # 36 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Cornell (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 60.33 ± 28.53 | # 33 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Deezer-Europe | MixHop | 1:1 Accuracy | 66.80±0.58 | # 11 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Film(48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 32.22 ± 2.34 | # 23 | |
| Node Classification | Film (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 33.13 ± 2.40 | # 30 | |
| Node Classification | genius | MixHop | Accuracy | 90.58 ± 0.16 | # 10 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | genius | MixHop | 1:1 Accuracy | 90.58 ± 0.16 | # 12 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Penn94 | MixHop | 1:1 Accuracy | 83.47 ± 0.71 | # 8 | |
| Node Classification | Penn94 | MixHop | Accuracy | 83.47 ± 0.71 | # 10 | |
| Node Classification | Pubmed | MixHop | Accuracy | 80.8% | # 31 | |
| Training Split | 20 per node | # 1 | ||||
| Validation | YES | # 1 | ||||
| Node Classification | PubMed (48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 85.31 ± 0.61 | # 25 | |
| Node Classification | PubMed (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 87.04 ± 4.10 | # 28 | |
| Node Classification | Squirrel | MixHop | Accuracy | 43.80 ± 1.48 | # 42 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Squirrel (48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 43.80 ± 1.48 | # 22 | |
| Node Classification | Squirrel (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 24.55 ± 2.60 | # 36 | |
| Node Classification | Texas | MixHop | Accuracy | 77.84 ± 7.73 | # 44 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Texas (48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 77.84 ± 7.73 | # 20 | |
| Node Classification | Texas (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 76.39 ± 7.66 | # 33 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Texas(60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 76.39 ± 7.66 | # 30 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | twitch-gamers | MixHop | 1:1 Accuracy | 65.64 ± 0.27 | # 10 | |
| Node Classification | Wisconsin | MixHop | Accuracy | 75.88 ± 4.90 | # 46 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Wisconsin (48%/32%/20% fixed splits) | MixHop | 1:1 Accuracy | 75.88 ± 4.90 | # 22 | |
| Node Classification | Wisconsin (60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 77.25 ± 7.80 | # 25 | |
| Node Classification on Non-Homophilic (Heterophilic) Graphs | Wisconsin(60%/20%/20% random splits) | MixHop | 1:1 Accuracy | 77.25 ± 7.80 | # 22 |