Robust Segmentation of Brain MRI in the Wild with Hierarchical CNNs and no Retraining
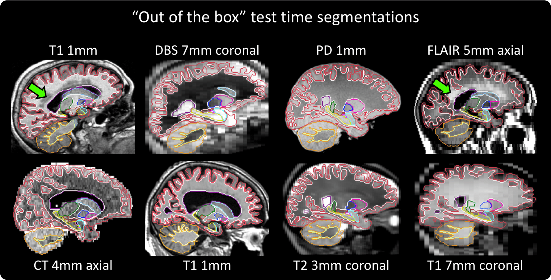
Retrospective analysis of brain MRI scans acquired in the clinic has the potential to enable neuroimaging studies with sample sizes much larger than those found in research datasets. However, analysing such clinical images "in the wild" is challenging, since subjects are scanned with highly variable protocols (MR contrast, resolution, orientation, etc.). Nevertheless, recent advances in convolutional neural networks (CNNs) and domain randomisation for image segmentation, best represented by the publicly available method SynthSeg, may enable morphometry of clinical MRI at scale. In this work, we first evaluate SynthSeg on an uncurated, heterogeneous dataset of more than 10,000 scans acquired at Massachusetts General Hospital. We show that SynthSeg is generally robust, but frequently falters on scans with low signal-to-noise ratio or poor tissue contrast. Next, we propose SynthSeg+, a novel method that greatly mitigates these problems using a hierarchy of conditional segmentation and denoising CNNs. We show that this method is considerably more robust than SynthSeg, while also outperforming cascaded networks and state-of-the-art segmentation denoising methods. Finally, we apply our approach to a proof-of-concept volumetric study of ageing, where it closely replicates atrophy patterns observed in research studies conducted on high-quality, 1mm, T1-weighted scans. The code and trained model are publicly available at https://github.com/BBillot/SynthSeg.
PDF Abstract