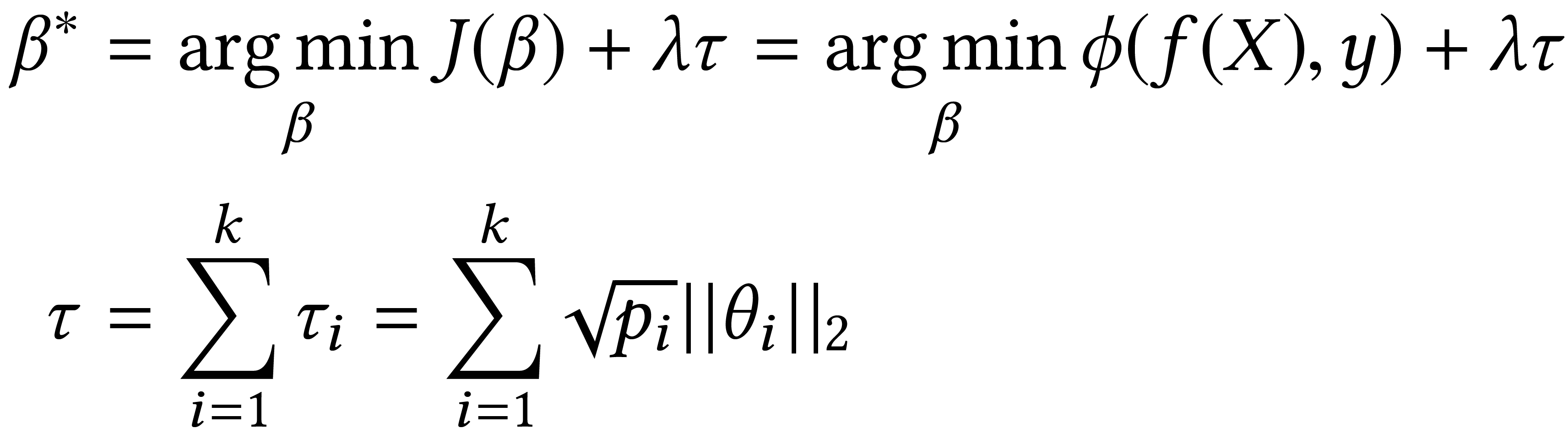Sparsely Grouped Input Variables for Neural Networks
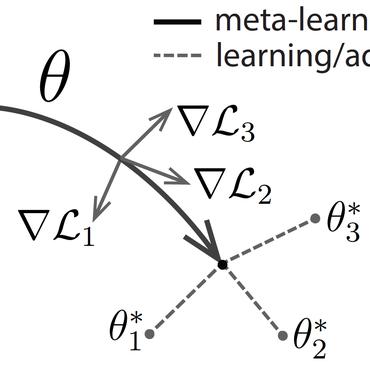
In genomic analysis, biomarker discovery, image recognition, and other systems involving machine learning, input variables can often be organized into different groups by their source or semantic category. Eliminating some groups of variables can expedite the process of data acquisition and avoid over-fitting. Researchers have used the group lasso to ensure group sparsity in linear models and have extended it to create compact neural networks in meta-learning. Different from previous studies, we use multi-layer non-linear neural networks to find sparse groups for input variables. We propose a new loss function to regularize parameters for grouped input variables, design a new optimization algorithm for this loss function, and test these methods in three real-world settings. We achieve group sparsity for three datasets, maintaining satisfying results while excluding one nucleotide position from an RNA splicing experiment, excluding 89.9% of stimuli from an eye-tracking experiment, and excluding 60% of image rows from an experiment on the MNIST dataset.
PDF Abstract