Gene set proximity analysis: expanding gene set enrichment analysis through learned geometric embeddings
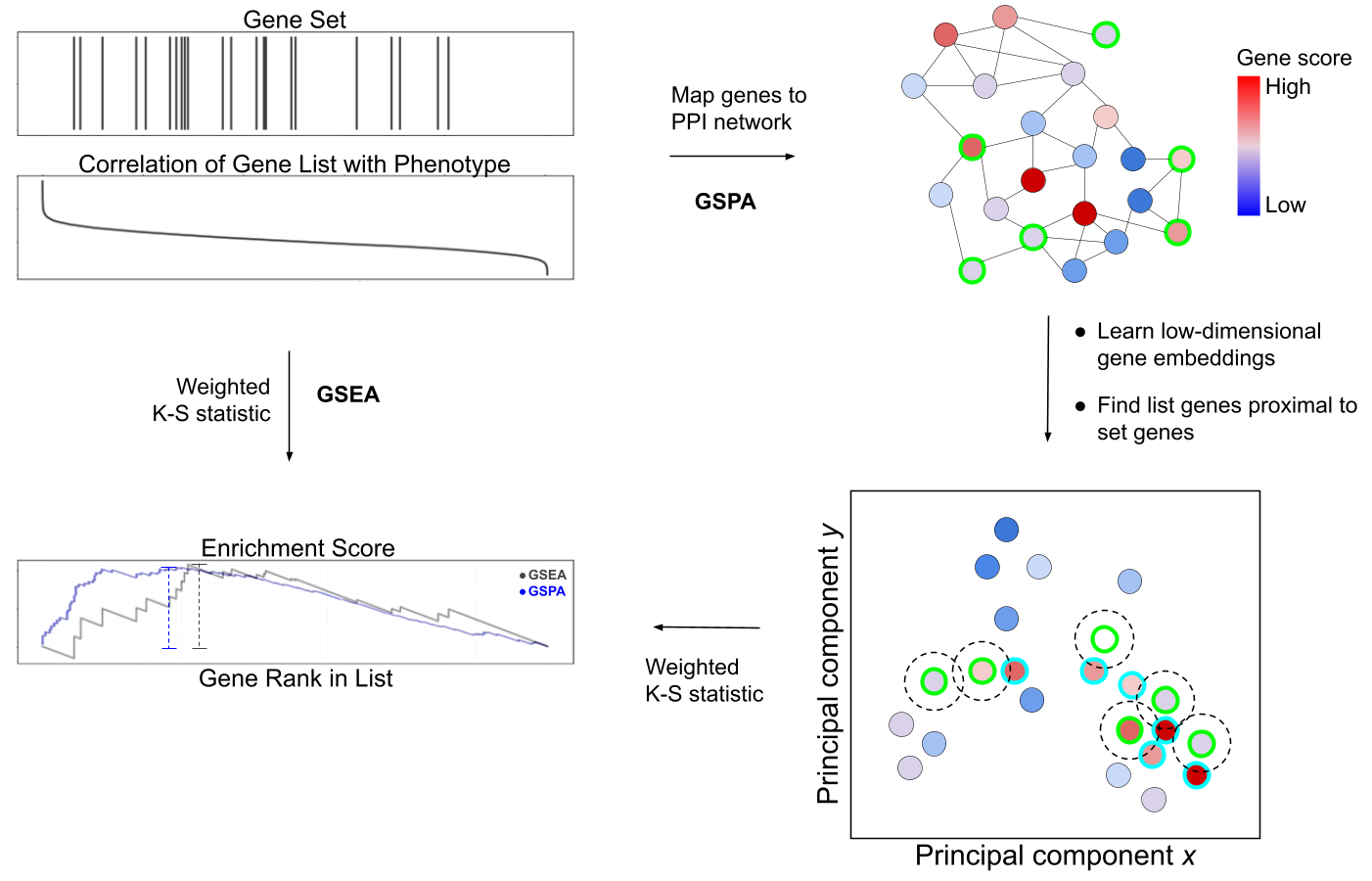
Gene set analysis methods rely on knowledge-based representations of genetic interactions in the form of both gene set collections and protein-protein interaction (PPI) networks. Explicit representations of genetic interactions often fail to capture complex interdependencies among genes, limiting the analytic power of such methods. Here we propose an extension of gene set enrichment analysis to a latent feature space reflecting PPI network topology, called gene set proximity analysis (GSPA). Compared with existing methods, GSPA provides improved ability to identify disease-associated pathways in disease-matched gene expression datasets, while improving reproducibility of enrichment statistics for similar gene sets. GSPA is statistically straightforward, reducing to classical gene set enrichment through a single user-defined parameter. We apply our method to identify novel drug associations with SARS-CoV-2 viral entry. Finally, we validate our drug association predictions through retrospective clinical analysis of claims data from 8 million patients, supporting a role for gabapentin as a risk factor and metformin as a protective factor for COVID-19 hospitalization.
PDF Abstract