Group SELFIES: A Robust Fragment-Based Molecular String Representation
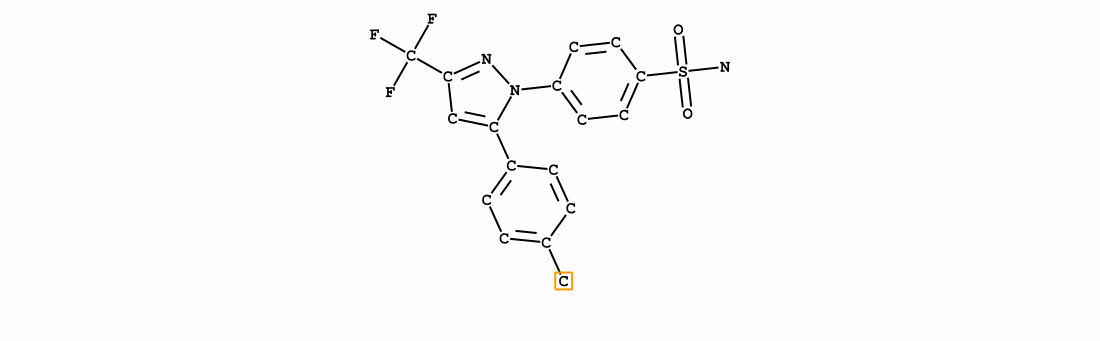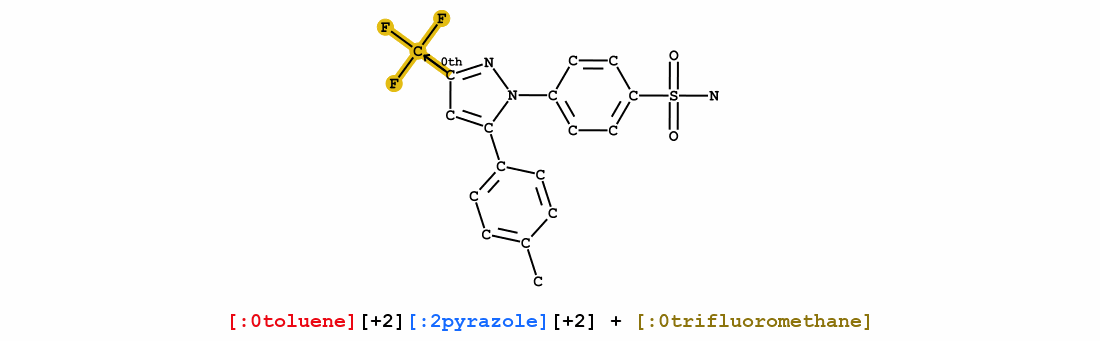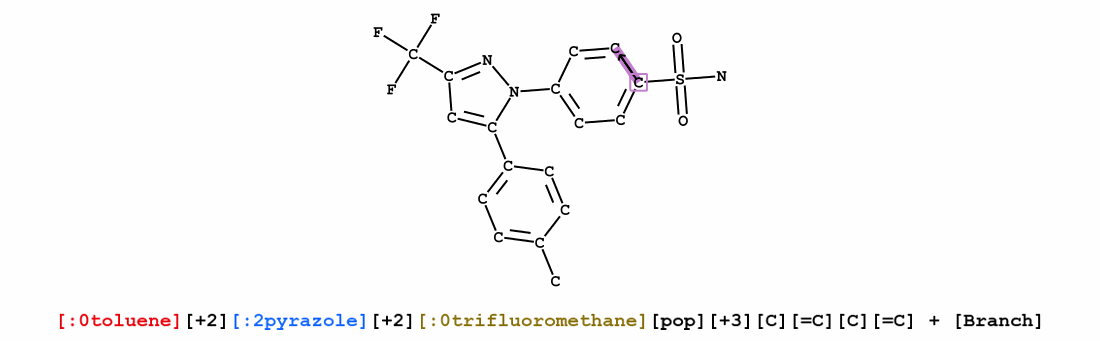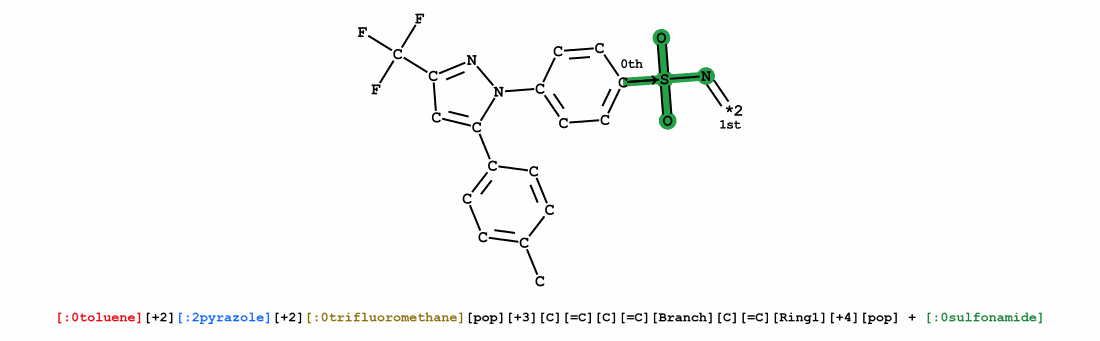
We introduce Group SELFIES, a molecular string representation that leverages group tokens to represent functional groups or entire substructures while maintaining chemical robustness guarantees. Molecular string representations, such as SMILES and SELFIES, serve as the basis for molecular generation and optimization in chemical language models, deep generative models, and evolutionary methods. While SMILES and SELFIES leverage atomic representations, Group SELFIES builds on top of the chemical robustness guarantees of SELFIES by enabling group tokens, thereby creating additional flexibility to the representation. Moreover, the group tokens in Group SELFIES can take advantage of inductive biases of molecular fragments that capture meaningful chemical motifs. The advantages of capturing chemical motifs and flexibility are demonstrated in our experiments, which show that Group SELFIES improves distribution learning of common molecular datasets. Further experiments also show that random sampling of Group SELFIES strings improves the quality of generated molecules compared to regular SELFIES strings. Our open-source implementation of Group SELFIES is available online, which we hope will aid future research in molecular generation and optimization.
PDF Abstract