Learning graph representations of biochemical networks and its application to enzymatic link prediction
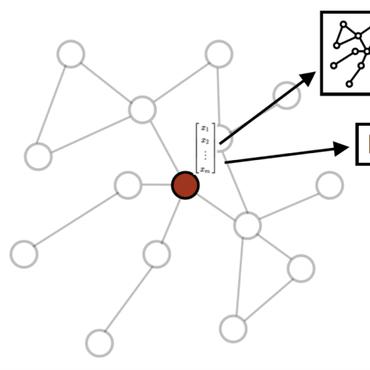
The complete characterization of enzymatic activities between molecules remains incomplete, hindering biological engineering and limiting biological discovery. We develop in this work a technique, Enzymatic Link Prediction (ELP), for predicting the likelihood of an enzymatic transformation between two molecules. ELP models enzymatic reactions catalogued in the KEGG database as a graph. ELP is innovative over prior works in using graph embedding to learn molecular representations that capture not only molecular and enzymatic attributes but also graph connectivity. We explore both transductive (test nodes included in the training graph) and inductive (test nodes not part of the training graph) learning models. We show that ELP achieves high AUC when learning node embeddings using both graph connectivity and node attributes. Further, we show that graph embedding for predicting enzymatic links improves link prediction by 24% over fingerprint-similarity-based approaches. To emphasize the importance of graph embedding in the context of biochemical networks, we illustrate how graph embedding can also guide visualization. The code and datasets are available through https://github.com/HassounLab/ELP.
PDF Abstract