Sub-word information in pre-trained biomedical word representations: evaluation and hyper-parameter optimization
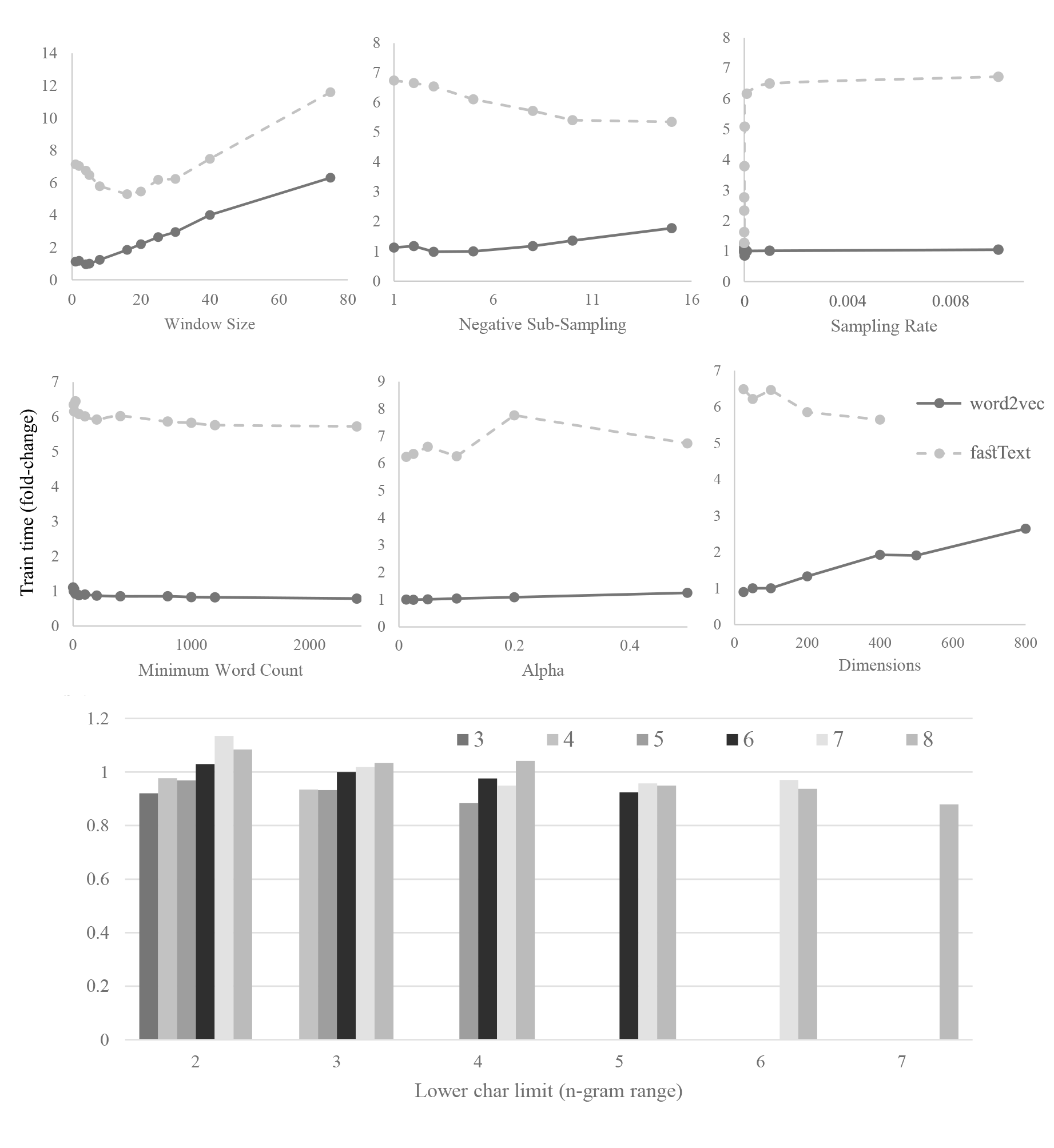
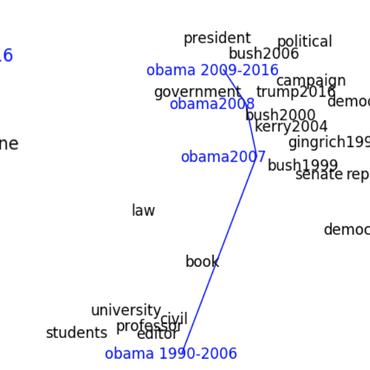
Word2vec embeddings are limited to computing vectors for in-vocabulary terms and do not take into account sub-word information. Character-based representations, such as fastText, mitigate such limitations. We optimize and compare these representations for the biomedical domain. fastText was found to consistently outperform word2vec in named entity recognition tasks for entities such as chemicals and genes. This is likely due to gained information from computed out-of-vocabulary term vectors, as well as the word compositionality of such entities. Contrastingly, performance varied on intrinsic datasets. Optimal hyper-parameters were intrinsic dataset-dependent, likely due to differences in term types distributions. This indicates embeddings should be chosen based on the task at hand. We therefore provide a number of optimized hyper-parameter sets and pre-trained word2vec and fastText models, available on \url{https://github.com/dterg/bionlp-embed}.
PDF Abstract