Transfer Learning in Biomedical Natural Language Processing: An Evaluation of BERT and ELMo on Ten Benchmarking Datasets
Code
Methods
No methods listed for this paper. Add
relevant methods here















 GLUE
GLUE
 MIMIC-III
MIMIC-III
 BC5CDR
BC5CDR
 DDI
DDI
 BIOSSES
BIOSSES
 HOC
HOC
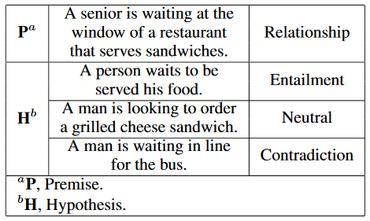
 MedNLI
MedNLI
