Deep neural networks with controlled variable selection for the identification of putative causal genetic variants
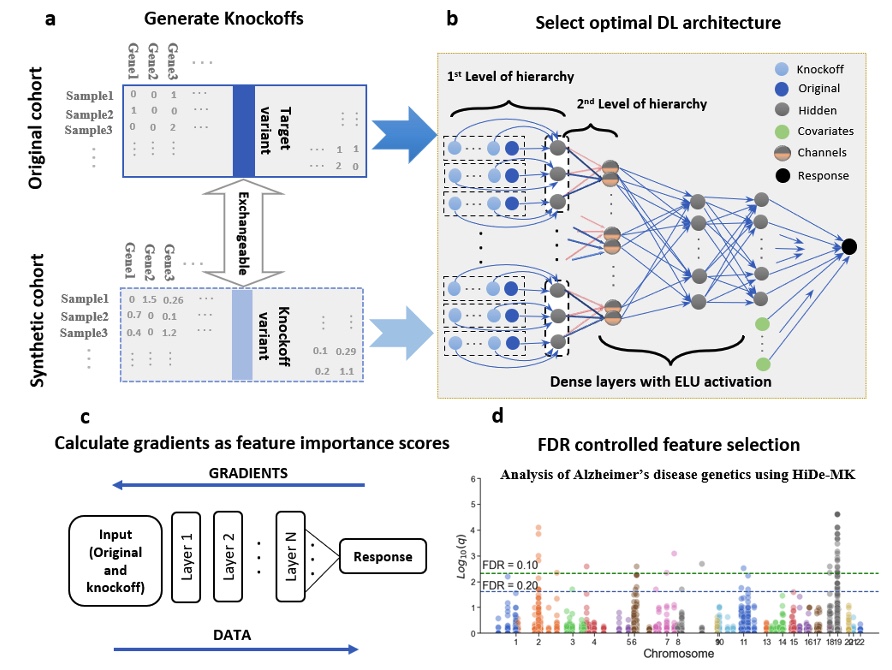
Deep neural networks (DNN) have been used successfully in many scientific problems for their high prediction accuracy, but their application to genetic studies remains challenging due to their poor interpretability. In this paper, we consider the problem of scalable, robust variable selection in DNN for the identification of putative causal genetic variants in genome sequencing studies. We identified a pronounced randomness in feature selection in DNN due to its stochastic nature, which may hinder interpretability and give rise to misleading results. We propose an interpretable neural network model, stabilized using ensembling, with controlled variable selection for genetic studies. The merit of the proposed method includes: (1) flexible modelling of the non-linear effect of genetic variants to improve statistical power; (2) multiple knockoffs in the input layer to rigorously control false discovery rate; (3) hierarchical layers to substantially reduce the number of weight parameters and activations to improve computational efficiency; (4) de-randomized feature selection to stabilize identified signals. We evaluated the proposed method in extensive simulation studies and applied it to the analysis of Alzheimer disease genetics. We showed that the proposed method, when compared to conventional linear and nonlinear methods, can lead to substantially more discoveries.
PDF Abstract