High Fidelity Deep Learning-based MRI Reconstruction with Instance-wise Discriminative Feature Matching Loss
Purpose: To improve reconstruction fidelity of fine structures and textures in deep learning (DL) based reconstructions. Methods: A novel patch-based Unsupervised Feature Loss (UFLoss) is proposed and incorporated into the training of DL-based reconstruction frameworks in order to preserve perceptual similarity and high-order statistics. The UFLoss provides instance-level discrimination by mapping similar instances to similar low-dimensional feature vectors and is trained without any human annotation. By adding an additional loss function on the low-dimensional feature space during training, the reconstruction frameworks from under-sampled or corrupted data can reproduce more realistic images that are closer to the original with finer textures, sharper edges, and improved overall image quality. The performance of the proposed UFLoss is demonstrated on unrolled networks for accelerated 2D and 3D knee MRI reconstruction with retrospective under-sampling. Quantitative metrics including NRMSE, SSIM, and our proposed UFLoss were used to evaluate the performance of the proposed method and compare it with others. Results: In-vivo experiments indicate that adding the UFLoss encourages sharper edges and more faithful contrasts compared to traditional and learning-based methods with pure l2 loss. More detailed textures can be seen in both 2D and 3D knee MR images. Quantitative results indicate that reconstruction with UFLoss can provide comparable NRMSE and a higher SSIM while achieving a much lower UFLoss value. Conclusion: We present UFLoss, a patch-based unsupervised learned feature loss, which allows the training of DL-based reconstruction to obtain more detailed texture, finer features, and sharper edges with higher overall image quality under DL-based reconstruction frameworks.
PDF Abstract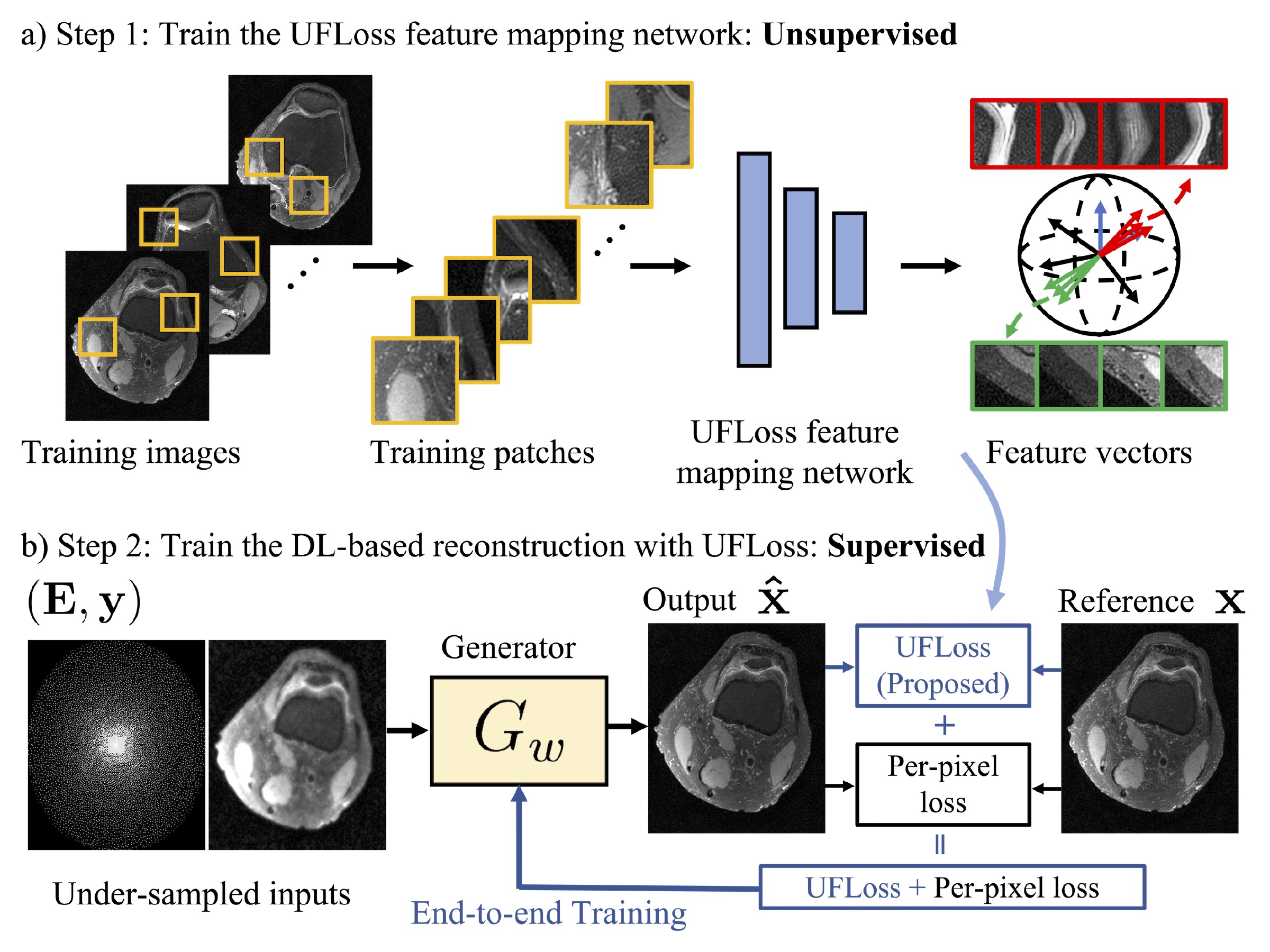


 Perceptual Similarity
Perceptual Similarity
 fastMRI
fastMRI