Search Results for author: Kaiyuan Gao
Found 10 papers, 8 papers with code
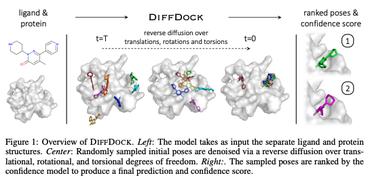
FABind+: Enhancing Molecular Docking through Improved Pocket Prediction and Pose Generation
1 code implementation • 29 Mar 2024 • Kaiyuan Gao, Qizhi Pei, Jinhua Zhu, Kun He, Lijun Wu
Molecular docking is a pivotal process in drug discovery.
Leveraging Biomolecule and Natural Language through Multi-Modal Learning: A Survey
2 code implementations • 3 Mar 2024 • Qizhi Pei, Lijun Wu, Kaiyuan Gao, Jinhua Zhu, Yue Wang, Zun Wang, Tao Qin, Rui Yan
The integration of biomolecular modeling with natural language (BL) has emerged as a promising interdisciplinary area at the intersection of artificial intelligence, chemistry and biology.
BioT5+: Towards Generalized Biological Understanding with IUPAC Integration and Multi-task Tuning
1 code implementation • 27 Feb 2024 • Qizhi Pei, Lijun Wu, Kaiyuan Gao, Xiaozhuan Liang, Yin Fang, Jinhua Zhu, Shufang Xie, Tao Qin, Rui Yan
However, previous efforts like BioT5 faced challenges in generalizing across diverse tasks and lacked a nuanced understanding of molecular structures, particularly in their textual representations (e. g., IUPAC).
 Ranked #1 on
Molecule Captioning
on ChEBI-20
Ranked #1 on
Molecule Captioning
on ChEBI-20
BioT5: Enriching Cross-modal Integration in Biology with Chemical Knowledge and Natural Language Associations
1 code implementation • 11 Oct 2023 • Qizhi Pei, Wei zhang, Jinhua Zhu, Kehan Wu, Kaiyuan Gao, Lijun Wu, Yingce Xia, Rui Yan
Recent advancements in biological research leverage the integration of molecules, proteins, and natural language to enhance drug discovery.
 Ranked #2 on
Text-based de novo Molecule Generation
on ChEBI-20
Ranked #2 on
Text-based de novo Molecule Generation
on ChEBI-20
FABind: Fast and Accurate Protein-Ligand Binding
1 code implementation • NeurIPS 2023 • Qizhi Pei, Kaiyuan Gao, Lijun Wu, Jinhua Zhu, Yingce Xia, Shufang Xie, Tao Qin, Kun He, Tie-Yan Liu, Rui Yan
In this work, we propose $\mathbf{FABind}$, an end-to-end model that combines pocket prediction and docking to achieve accurate and fast protein-ligand binding.
Examining User-Friendly and Open-Sourced Large GPT Models: A Survey on Language, Multimodal, and Scientific GPT Models
1 code implementation • 27 Aug 2023 • Kaiyuan Gao, Sunan He, Zhenyu He, Jiacheng Lin, Qizhi Pei, Jie Shao, Wei zhang
Generative pre-trained transformer (GPT) models have revolutionized the field of natural language processing (NLP) with remarkable performance in various tasks and also extend their power to multimodal domains.
Tokenized Graph Transformer with Neighborhood Augmentation for Node Classification in Large Graphs
no code implementations • 22 May 2023 • Jinsong Chen, Chang Liu, Kaiyuan Gao, Gaichao Li, Kun He
Graph Transformers, emerging as a new architecture for graph representation learning, suffer from the quadratic complexity on the number of nodes when handling large graphs.
Incorporating Pre-training Paradigm for Antibody Sequence-Structure Co-design
no code implementations • 26 Oct 2022 • Kaiyuan Gao, Lijun Wu, Jinhua Zhu, Tianbo Peng, Yingce Xia, Liang He, Shufang Xie, Tao Qin, Haiguang Liu, Kun He, Tie-Yan Liu
Specifically, we first pre-train an antibody language model based on the sequence data, then propose a one-shot way for sequence and structure generation of CDR to avoid the heavy cost and error propagation from an autoregressive manner, and finally leverage the pre-trained antibody model for the antigen-specific antibody generation model with some carefully designed modules.
NAGphormer: A Tokenized Graph Transformer for Node Classification in Large Graphs
1 code implementation • 10 Jun 2022 • Jinsong Chen, Kaiyuan Gao, Gaichao Li, Kun He
In this work, we observe that existing graph Transformers treat nodes as independent tokens and construct a single long sequence composed of all node tokens so as to train the Transformer model, causing it hard to scale to large graphs due to the quadratic complexity on the number of nodes for the self-attention computation.
Revisiting Language Encoding in Learning Multilingual Representations
1 code implementation • 16 Feb 2021 • Shengjie Luo, Kaiyuan Gao, Shuxin Zheng, Guolin Ke, Di He, LiWei Wang, Tie-Yan Liu
The language embedding can be either added to the word embedding or attached at the beginning of the sentence.