Adapting Segment Anything Model (SAM) through Prompt-based Learning for Enhanced Protein Identification in Cryo-EM Micrographs
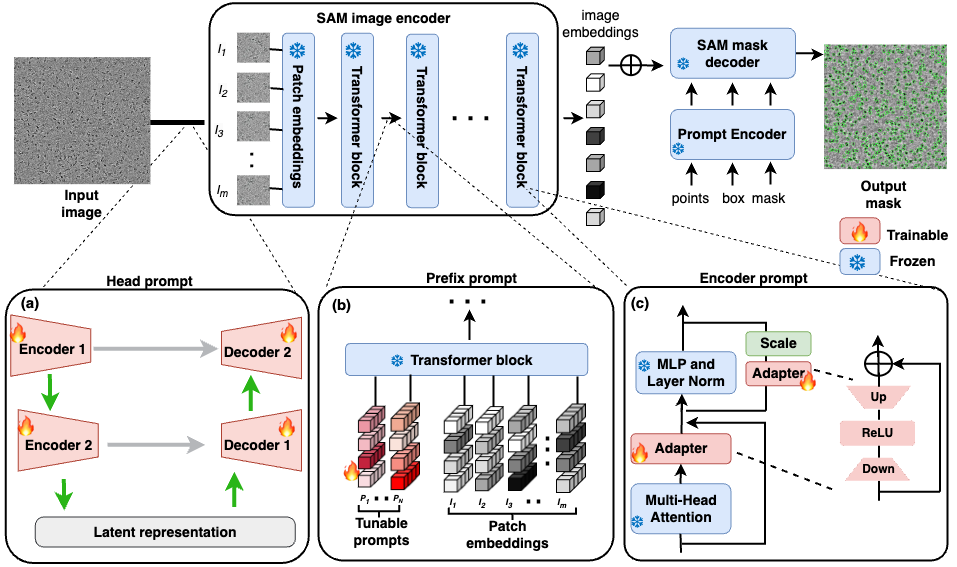
Cryo-electron microscopy (cryo-EM) remains pivotal in structural biology, yet the task of protein particle picking, integral for 3D protein structure construction, is laden with manual inefficiencies. While recent AI tools such as Topaz and crYOLO are advancing the field, they do not fully address the challenges of cryo-EM images, including low contrast, complex shapes, and heterogeneous conformations. This study explored prompt-based learning to adapt the state-of-the-art image segmentation foundation model Segment Anything Model (SAM) for cryo-EM. This focus was driven by the desire to optimize model performance with a small number of labeled data without altering pre-trained parameters, aiming for a balance between adaptability and foundational knowledge retention. Through trials with three prompt-based learning strategies, namely head prompt, prefix prompt, and encoder prompt, we observed enhanced performance and reduced computational requirements compared to the fine-tuning approach. This work not only highlights the potential of prompting SAM in protein identification from cryo-EM micrographs but also suggests its broader promise in biomedical image segmentation and object detection.
PDF Abstract