Medical Image Registration
77 papers with code • 4 benchmarks • 8 datasets
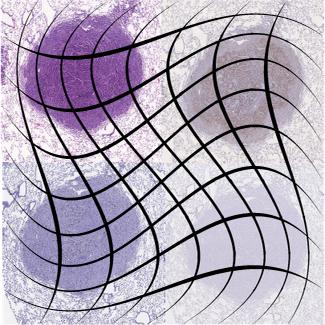
Image registration, also known as image fusion or image matching, is the process of aligning two or more images based on image appearances. Medical Image Registration seeks to find an optimal spatial transformation that best aligns the underlying anatomical structures. Medical Image Registration is used in many clinical applications such as image guidance, motion tracking, segmentation, dose accumulation, image reconstruction and so on. Medical Image Registration is a broad topic which can be grouped from various perspectives. From input image point of view, registration methods can be divided into unimodal, multimodal, interpatient, intra-patient (e.g. same- or different-day) registration. From deformation model point of view, registration methods can be divided in to rigid, affine and deformable methods. From region of interest (ROI) perspective, registration methods can be grouped according to anatomical sites such as brain, lung registration and so on. From image pair dimension perspective, registration methods can be divided into 3D to 3D, 3D to 2D and 2D to 2D/3D.
Source: Deep Learning in Medical Image Registration: A Review
Libraries
Use these libraries to find Medical Image Registration models and implementationsDatasets
Most implemented papers
VoxelMorph: A Learning Framework for Deformable Medical Image Registration
In contrast to this approach, and building on recent learning-based methods, we formulate registration as a function that maps an input image pair to a deformation field that aligns these images.
Unsupervised 3D End-to-End Medical Image Registration with Volume Tweening Network
3D medical image registration is of great clinical importance.
Recursive Cascaded Networks for Unsupervised Medical Image Registration
We present recursive cascaded networks, a general architecture that enables learning deep cascades, for deformable image registration.
An Unsupervised Learning Model for Deformable Medical Image Registration
We define registration as a parametric function, and optimize its parameters given a set of images from a collection of interest.
AirLab: Autograd Image Registration Laboratory
With the "Autograd Image Registration Laboratory" (AIRLab), we introduce an open laboratory for image registration tasks, where the analytic gradients of the objective function are computed automatically and the device where the computations are performed, on a CPU or a GPU, is transparent.
Fast and Robust Symmetric Image Registration Based on Distances Combining Intensity and Spatial Information
The method exhibits greater robustness and higher accuracy than similarity measures in common use, when inserted into a standard gradient-based registration framework available as part of the open source Insight Segmentation and Registration Toolkit (ITK).
Networks for Joint Affine and Non-parametric Image Registration
In contrast to existing approaches, our framework combines two registration methods: an affine registration and a vector momentum-parameterized stationary velocity field (vSVF) model.
Dual-Stream Pyramid Registration Network
We propose a Dual-Stream Pyramid Registration Network (referred as Dual-PRNet) for unsupervised 3D medical image registration.
Enhancing Medical Image Registration via Appearance Adjustment Networks
In this study, we propose an Appearance Adjustment Network (AAN) to enhance the adaptability of DLRs to appearance variations.
Generative Adversarial Registration for Improved Conditional Deformable Templates
Deformable templates are essential to large-scale medical image registration, segmentation, and population analysis.




 PPMI
PPMI
 Learn2Reg
Learn2Reg
 OASIS
OASIS
 IXI
IXI
 SR-Reg
SR-Reg
 Full-Spectral Autofluorescence Lifetime Microscopic Images
Full-Spectral Autofluorescence Lifetime Microscopic Images
 BreastDICOM4
BreastDICOM4