Search Results for author: Nils D. Forkert
Found 10 papers, 1 papers with code
Towards objective and systematic evaluation of bias in medical imaging AI
1 code implementation • 3 Nov 2023 • Emma A. M. Stanley, Raissa Souza, Anthony Winder, Vedant Gulve, Kimberly Amador, Matthias Wilms, Nils D. Forkert
In this article, we introduce a novel analysis framework for systematically and objectively investigating the impact of biases in medical images on AI models.
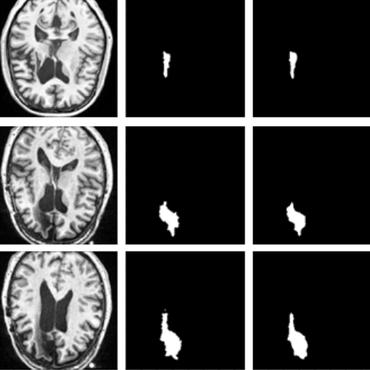
Weakly Supervised Medical Image Segmentation With Soft Labels and Noise Robust Loss
no code implementations • 16 Sep 2022 • Banafshe Felfeliyan, Abhilash Hareendranathan, Gregor Kuntze, Stephanie Wichuk, Nils D. Forkert, Jacob L. Jaremko, Janet L. Ronsky
The aim of this paper was to develop and evaluate a method to generate probabilistic labels based on multi-rater annotations and anatomical knowledge of the lesion features in MRI and a method to train segmentation models using probabilistic labels using normalized active-passive loss as a "noise-tolerant loss" function.
Self-Supervised-RCNN for Medical Image Segmentation with Limited Data Annotation
no code implementations • 17 Jul 2022 • Banafshe Felfeliyan, Abhilash Hareendranathan, Gregor Kuntze, David Cornell, Nils D. Forkert, Jacob L. Jaremko, Janet L. Ronsky
The effectiveness of the proposed method for segmentation tasks in different pre-training and fine-tuning scenarios is evaluated based on the Osteoarthritis Initiative dataset.
Bone Adaptation as a Geometric Flow
no code implementations • 9 Nov 2021 • Bryce A. Besler, Tannis D. Kemp, Nils D. Forkert, Steven K. Boyd
Interestingly, the parameters of the model can be defined in such a way that the flow is volume-preserving.
Constructing High-Order Signed Distance Maps from Computed Tomography Data with Application to Bone Morphometry
no code implementations • 2 Nov 2021 • Bryce A. Besler, Tannis D. Kemp, Nils D. Forkert, Steven K. Boyd
The narrowband is solved using a closest point algorithm extended for implicit embeddings that are not a signed distance field.
High-Order Signed Distance Transform of Sampled Signals
no code implementations • 26 Oct 2021 • Bryce A. Besler, Tannis D. Kemp, Nils D. Forkert, Steven K. Boyd
Such a transform is an improvement to the classic notion of an exact signed distance transform because it does not exhibit artifacts of quantization.
Local Morphometry of Closed, Implicit Surfaces
no code implementations • 29 Jul 2021 • Bryce A Besler, Tannis D. Kemp, Andrew S. Michalski, Nils D. Forkert, Steven K. Boyd
The proposed method enables an improved local and global evaluation of curvature for purposes of morphometry on closed, implicit surfaces.
Bidirectional Modeling and Analysis of Brain Aging with Normalizing Flows
no code implementations • 26 Nov 2020 • Matthias Wilms, Jordan J. Bannister, Pauline Mouches, M. Ethan MacDonald, Deepthi Rajashekar, Sönke Langner, Nils D. Forkert
Brain aging is a widely studied longitudinal process throughout which the brain undergoes considerable morphological changes and various machine learning approaches have been proposed to analyze it.
Automatic segmentation of stroke lesions in non-contrast computed tomography with convolutional neural networks
no code implementations • MIDL 2019 • Anup Tuladhar, Serena Schimert, Deepthi Rajashekar, Helge C. Kniep, Jens Fiehler, Nils D. Forkert
A total of 291 multi-center clinical NCCT datasets were used: 204 for CNN training, 48 for validation and developing post-processing methods, and 39 for testing.
DeepVesselNet: Vessel Segmentation, Centerline Prediction, and Bifurcation Detection in 3-D Angiographic Volumes
no code implementations • 25 Mar 2018 • Giles Tetteh, Velizar Efremov, Nils D. Forkert, Matthias Schneider, Jan Kirschke, Bruno Weber, Claus Zimmer, Marie Piraud, Bjoern H. Menze
Our experiments show that, by replacing 3-D filters with cross-hair filters in our network, we achieve over 23% improvement in speed, lower memory footprint, lower network complexity which prevents overfitting and comparable accuracy (with a Cox-Wilcoxon paired sample significance test p-value of 0. 07 when compared to full 3-D filters).