Learning from partially labeled data for multi-organ and tumor segmentation
Medical image benchmarks for the segmentation of organs and tumors suffer from the partially labeling issue due to its intensive cost of labor and expertise. Current mainstream approaches follow the practice of one network solving one task. With this pipeline, not only the performance is limited by the typically small dataset of a single task, but also the computation cost linearly increases with the number of tasks. To address this, we propose a Transformer based dynamic on-demand network (TransDoDNet) that learns to segment organs and tumors on multiple partially labeled datasets. Specifically, TransDoDNet has a hybrid backbone that is composed of the convolutional neural network and Transformer. A dynamic head enables the network to accomplish multiple segmentation tasks flexibly. Unlike existing approaches that fix kernels after training, the kernels in the dynamic head are generated adaptively by the Transformer, which employs the self-attention mechanism to model long-range organ-wise dependencies and decodes the organ embedding that can represent each organ. We create a large-scale partially labeled Multi-Organ and Tumor Segmentation benchmark, termed MOTS, and demonstrate the superior performance of our TransDoDNet over other competitors on seven organ and tumor segmentation tasks. This study also provides a general 3D medical image segmentation model, which has been pre-trained on the large-scale MOTS benchmark and has demonstrated advanced performance over BYOL, the current predominant self-supervised learning method. Code will be available at \url{https://git.io/DoDNet}.
PDF Abstract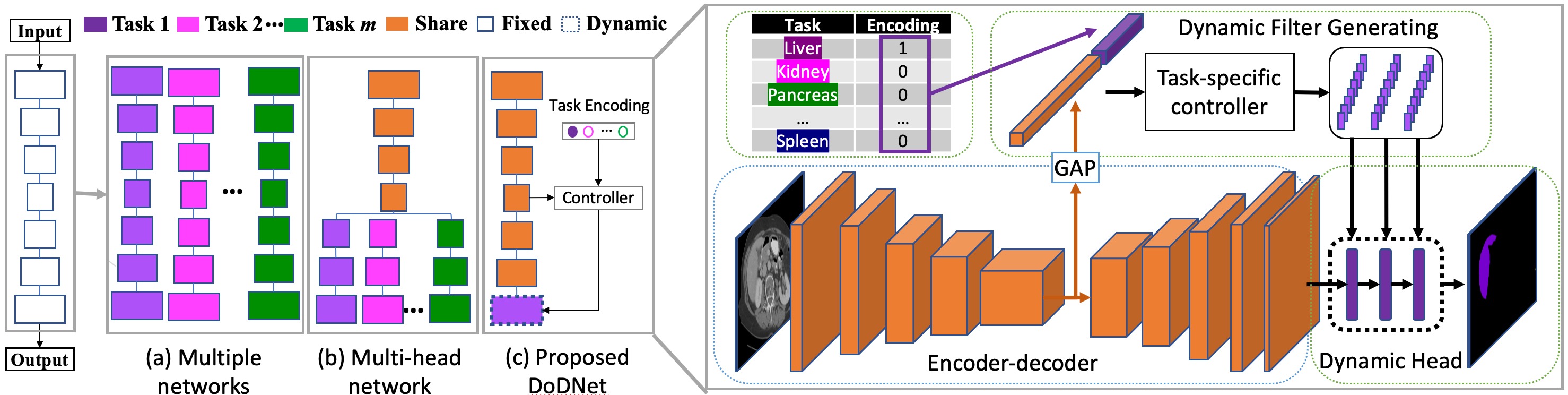






 Cityscapes
Cityscapes