Search Results for author: Wenjia Bai
Found 69 papers, 28 papers with code
A Foundation Model for Brain Lesion Segmentation with Mixture of Modality Experts
no code implementations • 16 May 2024 • Xinru Zhang, Ni Ou, Berke Doga Basaran, Marco Visentin, Mengyun Qiao, Renyang Gu, Cheng Ouyang, Yaou Liu, Paul M. Matthew, Chuyang Ye, Wenjia Bai
In this work, we propose a universal foundation model for 3D brain lesion segmentation, which can automatically segment different types of brain lesions for input data of various imaging modalities.
The state-of-the-art in Cardiac MRI Reconstruction: Results of the CMRxRecon Challenge in MICCAI 2023
no code implementations • 1 Apr 2024 • Jun Lyu, Chen Qin, Shuo Wang, Fanwen Wang, Yan Li, Zi Wang, Kunyuan Guo, Cheng Ouyang, Michael Tänzer, Meng Liu, Longyu Sun, Mengting Sun, Qin Li, Zhang Shi, Sha Hua, Hao Li, Zhensen Chen, Zhenlin Zhang, Bingyu Xin, Dimitris N. Metaxas, George Yiasemis, Jonas Teuwen, Liping Zhang, Weitian Chen, Yidong Zhao, Qian Tao, Yanwei Pang, Xiaohan Liu, Artem Razumov, Dmitry V. Dylov, Quan Dou, Kang Yan, Yuyang Xue, Yuning Du, Julia Dietlmeier, Carles Garcia-Cabrera, Ziad Al-Haj Hemidi, Nora Vogt, Ziqiang Xu, Yajing Zhang, Ying-Hua Chu, Weibo Chen, Wenjia Bai, Xiahai Zhuang, Jing Qin, Lianmin Wu, Guang Yang, Xiaobo Qu, He Wang, Chengyan Wang
To address this issue, we organized the Cardiac MRI Reconstruction Challenge (CMRxRecon) in 2023, in collaboration with the 26th International Conference on MICCAI.
Zero-Shot ECG Classification with Multimodal Learning and Test-time Clinical Knowledge Enhancement
no code implementations • 11 Mar 2024 • Che Liu, Zhongwei Wan, Cheng Ouyang, Anand Shah, Wenjia Bai, Rossella Arcucci
Through multimodal learning on ECG records and associated reports, MERL is capable of performing zero-shot ECG classification with text prompts, eliminating the need for training data in downstream tasks.
G2D: From Global to Dense Radiography Representation Learning via Vision-Language Pre-training
no code implementations • 3 Dec 2023 • Che Liu, Cheng Ouyang, Sibo Cheng, Anand Shah, Wenjia Bai, Rossella Arcucci
G2D achieves superior performance across 6 medical imaging tasks and 25 diseases, particularly in semantic segmentation, which necessitates fine-grained, semantically-grounded image features.
T3D: Towards 3D Medical Image Understanding through Vision-Language Pre-training
no code implementations • 3 Dec 2023 • Che Liu, Cheng Ouyang, Yinda Chen, Cesar César Quilodrán-Casas, Lei Ma, Jie Fu, Yike Guo, Anand Shah, Wenjia Bai, Rossella Arcucci
This underlines T3D's potential in representation learning for 3D medical image analysis.
IMITATE: Clinical Prior Guided Hierarchical Vision-Language Pre-training
no code implementations • 11 Oct 2023 • Che Liu, Sibo Cheng, Miaojing Shi, Anand Shah, Wenjia Bai, Rossella Arcucci
The framework derives multi-level visual features from the chest X-ray (CXR) images and separately aligns these features with the descriptive and the conclusive text encoded in the hierarchical medical report.
Utilizing Synthetic Data for Medical Vision-Language Pre-training: Bypassing the Need for Real Images
1 code implementation • 10 Oct 2023 • Che Liu, Anand Shah, Wenjia Bai, Rossella Arcucci
The advent of text-guided generative models raises a compelling question: Can VLP be implemented solely with synthetic images generated from genuine radiology reports, thereby mitigating the need for extensively pairing and curating image-text datasets?
T1/T2 relaxation temporal modelling from accelerated acquisitions using a Latent Transformer
no code implementations • 28 Sep 2023 • Fanwen Wang, Michael Tanzer, Mengyun Qiao, Wenjia Bai, Daniel Rueckert, Guang Yang, Sonia Nielles-Vallespin
Quantitative cardiac magnetic resonance T1 and T2 mapping enable myocardial tissue characterisation but the lengthy scan times restrict their widespread clinical application.
DeepMesh: Mesh-based Cardiac Motion Tracking using Deep Learning
no code implementations • 25 Sep 2023 • Qingjie Meng, Wenjia Bai, Declan P O'Regan, and Daniel Rueckert
We propose a novel learning framework, DeepMesh, which propagates a template heart mesh to a subject space and estimates the 3D motion of the heart mesh from CMR images for individual subjects.
CMRxRecon: An open cardiac MRI dataset for the competition of accelerated image reconstruction
1 code implementation • 19 Sep 2023 • Chengyan Wang, Jun Lyu, Shuo Wang, Chen Qin, Kunyuan Guo, Xinyu Zhang, Xiaotong Yu, Yan Li, Fanwen Wang, Jianhua Jin, Zhang Shi, Ziqiang Xu, Yapeng Tian, Sha Hua, Zhensen Chen, Meng Liu, Mengting Sun, Xutong Kuang, Kang Wang, Haoran Wang, Hao Li, Yinghua Chu, Guang Yang, Wenjia Bai, Xiahai Zhuang, He Wang, Jing Qin, Xiaobo Qu
However, a limitation of CMR is its slow imaging speed, which causes patient discomfort and introduces artifacts in the images.
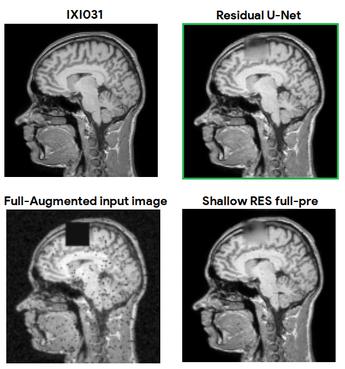
LesionMix: A Lesion-Level Data Augmentation Method for Medical Image Segmentation
1 code implementation • 17 Aug 2023 • Berke Doga Basaran, Weitong Zhang, Mengyun Qiao, Bernhard Kainz, Paul M. Matthews, Wenjia Bai
Data augmentation has become a de facto component of deep learning-based medical image segmentation methods.
Hierarchical Uncertainty Estimation for Medical Image Segmentation Networks
no code implementations • 16 Aug 2023 • Xinyu Bai, Wenjia Bai
Learning a medical image segmentation model is an inherently ambiguous task, as uncertainties exist in both images (noise) and manual annotations (human errors and bias) used for model training.
M-FLAG: Medical Vision-Language Pre-training with Frozen Language Models and Latent Space Geometry Optimization
1 code implementation • 17 Jul 2023 • Che Liu, Sibo Cheng, Chen Chen, Mengyun Qiao, Weitong Zhang, Anand Shah, Wenjia Bai, Rossella Arcucci
The proposed method, named Medical vision-language pre-training with Frozen language models and Latent spAce Geometry optimization (M-FLAG), leverages a frozen language model for training stability and efficiency and introduces a novel orthogonality loss to harmonize the latent space geometry.
CHeart: A Conditional Spatio-Temporal Generative Model for Cardiac Anatomy
1 code implementation • 30 Jan 2023 • Mengyun Qiao, Shuo Wang, Huaqi Qiu, Antonio de Marvao, Declan P. O'Regan, Daniel Rueckert, Wenjia Bai
Two key questions in cardiac image analysis are to assess the anatomy and motion of the heart from images; and to understand how they are associated with non-imaging clinical factors such as gender, age and diseases.
The Extreme Cardiac MRI Analysis Challenge under Respiratory Motion (CMRxMotion)
no code implementations • 12 Oct 2022 • Shuo Wang, Chen Qin, Chengyan Wang, Kang Wang, Haoran Wang, Chen Chen, Cheng Ouyang, Xutong Kuang, Chengliang Dai, Yuanhan Mo, Zhang Shi, Chenchen Dai, Xinrong Chen, He Wang, Wenjia Bai
The quality of cardiac magnetic resonance (CMR) imaging is susceptible to respiratory motion artifacts.
Mesh-based 3D Motion Tracking in Cardiac MRI using Deep Learning
no code implementations • 5 Sep 2022 • Qingjie Meng, Wenjia Bai, Tianrui Liu, Declan P O'Regan, Daniel Rueckert
By developing a differentiable mesh-to-image rasterizer, the method is able to leverage the anatomical shape information from 2D multi-view CMR images for 3D motion estimation.
Generative Modelling of the Ageing Heart with Cross-Sectional Imaging and Clinical Data
1 code implementation • 28 Aug 2022 • Mengyun Qiao, Berke Doga Basaran, Huaqi Qiu, Shuo Wang, Yi Guo, Yuanyuan Wang, Paul M. Matthews, Daniel Rueckert, Wenjia Bai
Understanding the morphological and functional changes of the heart during ageing is a key scientific question, the answer to which will help us define important risk factors of cardiovascular disease and monitor disease progression.
Improved post-hoc probability calibration for out-of-domain MRI segmentation
1 code implementation • 4 Aug 2022 • Cheng Ouyang, Shuo Wang, Chen Chen, Zeju Li, Wenjia Bai, Bernhard Kainz, Daniel Rueckert
In image segmentation, well-calibrated probabilities allow radiologists to identify regions where model-predicted segmentations are unreliable.
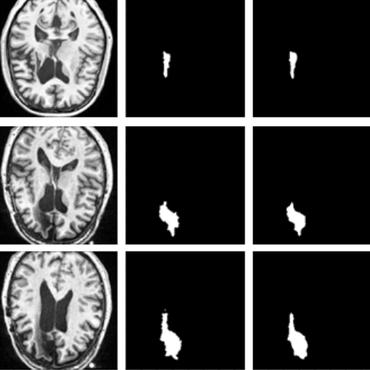
Subject-Specific Lesion Generation and Pseudo-Healthy Synthesis for Multiple Sclerosis Brain Images
1 code implementation • 3 Aug 2022 • Berke Doga Basaran, Mengyun Qiao, Paul M. Matthews, Wenjia Bai
In this work, we present a novel foreground-based generative method for modelling the local lesion characteristics that can both generate synthetic lesions on healthy images and synthesize subject-specific pseudo-healthy images from pathological images.
MulViMotion: Shape-aware 3D Myocardial Motion Tracking from Multi-View Cardiac MRI
no code implementations • 29 Jul 2022 • Qingjie Meng, Chen Qin, Wenjia Bai, Tianrui Liu, Antonio de Marvao, Declan P O'Regan, Daniel Rueckert
To address this problem, we propose a novel multi-view motion estimation network (MulViMotion), which integrates 2D cine CMR images acquired in short-axis and long-axis planes to learn a consistent 3D motion field of the heart.
Generative Myocardial Motion Tracking via Latent Space Exploration with Biomechanics-informed Prior
1 code implementation • 8 Jun 2022 • Chen Qin, Shuo Wang, Chen Chen, Wenjia Bai, Daniel Rueckert
In contrast to most existing approaches which impose explicit generic regularization such as smoothness, in this work we propose a novel method that can implicitly learn an application-specific biomechanics-informed prior and embed it into a neural network-parameterized transformation model.
MaxStyle: Adversarial Style Composition for Robust Medical Image Segmentation
1 code implementation • 2 Jun 2022 • Chen Chen, Zeju Li, Cheng Ouyang, Matt Sinclair, Wenjia Bai, Daniel Rueckert
We propose a novel data augmentation framework called MaxStyle, which maximizes the effectiveness of style augmentation for model OOD performance.
Suggestive Annotation of Brain MR Images with Gradient-guided Sampling
no code implementations • 2 Jun 2022 • Chengliang Dai, Shuo Wang, Yuanhan Mo, Elsa Angelini, Yike Guo, Wenjia Bai
We evaluate the framework on two different brain image analysis tasks, namely brain tumour segmentation and whole brain segmentation.
Memory-efficient Segmentation of High-resolution Volumetric MicroCT Images
1 code implementation • 31 May 2022 • YuAn Wang, Laura Blackie, Irene Miguel-Aliaga, Wenjia Bai
In this work, we propose a novel memory-efficient network architecture for 3D high-resolution image segmentation.
QU-BraTS: MICCAI BraTS 2020 Challenge on Quantifying Uncertainty in Brain Tumor Segmentation - Analysis of Ranking Scores and Benchmarking Results
1 code implementation • 19 Dec 2021 • Raghav Mehta, Angelos Filos, Ujjwal Baid, Chiharu Sako, Richard McKinley, Michael Rebsamen, Katrin Datwyler, Raphael Meier, Piotr Radojewski, Gowtham Krishnan Murugesan, Sahil Nalawade, Chandan Ganesh, Ben Wagner, Fang F. Yu, Baowei Fei, Ananth J. Madhuranthakam, Joseph A. Maldjian, Laura Daza, Catalina Gomez, Pablo Arbelaez, Chengliang Dai, Shuo Wang, Hadrien Reynaud, Yuan-han Mo, Elsa Angelini, Yike Guo, Wenjia Bai, Subhashis Banerjee, Lin-min Pei, Murat AK, Sarahi Rosas-Gonzalez, Ilyess Zemmoura, Clovis Tauber, Minh H. Vu, Tufve Nyholm, Tommy Lofstedt, Laura Mora Ballestar, Veronica Vilaplana, Hugh McHugh, Gonzalo Maso Talou, Alan Wang, Jay Patel, Ken Chang, Katharina Hoebel, Mishka Gidwani, Nishanth Arun, Sharut Gupta, Mehak Aggarwal, Praveer Singh, Elizabeth R. Gerstner, Jayashree Kalpathy-Cramer, Nicolas Boutry, Alexis Huard, Lasitha Vidyaratne, Md Monibor Rahman, Khan M. Iftekharuddin, Joseph Chazalon, Elodie Puybareau, Guillaume Tochon, Jun Ma, Mariano Cabezas, Xavier Llado, Arnau Oliver, Liliana Valencia, Sergi Valverde, Mehdi Amian, Mohammadreza Soltaninejad, Andriy Myronenko, Ali Hatamizadeh, Xue Feng, Quan Dou, Nicholas Tustison, Craig Meyer, Nisarg A. Shah, Sanjay Talbar, Marc-Andre Weber, Abhishek Mahajan, Andras Jakab, Roland Wiest, Hassan M. Fathallah-Shaykh, Arash Nazeri, Mikhail Milchenko1, Daniel Marcus, Aikaterini Kotrotsou, Rivka Colen, John Freymann, Justin Kirby, Christos Davatzikos, Bjoern Menze, Spyridon Bakas, Yarin Gal, Tal Arbel
In this study, we explore and evaluate a score developed during the BraTS 2019 and BraTS 2020 task on uncertainty quantification (QU-BraTS) and designed to assess and rank uncertainty estimates for brain tumor multi-compartment segmentation.
Causality-inspired Single-source Domain Generalization for Medical Image Segmentation
1 code implementation • 24 Nov 2021 • Cheng Ouyang, Chen Chen, Surui Li, Zeju Li, Chen Qin, Wenjia Bai, Daniel Rueckert
In this work, we investigate the single-source domain generalization problem: training a deep network that is robust to unseen domains, under the condition that training data is only available from one source domain, which is common in medical imaging applications.
DeepMCAT: Large-Scale Deep Clustering for Medical Image Categorization
no code implementations • 30 Sep 2021 • Turkay Kart, Wenjia Bai, Ben Glocker, Daniel Rueckert
In recent years, the research landscape of machine learning in medical imaging has changed drastically from supervised to semi-, weakly- or unsupervised methods.
Enhancing MR Image Segmentation with Realistic Adversarial Data Augmentation
1 code implementation • 7 Aug 2021 • Chen Chen, Chen Qin, Cheng Ouyang, Zeju Li, Shuo Wang, Huaqi Qiu, Liang Chen, Giacomo Tarroni, Wenjia Bai, Daniel Rueckert
The success of neural networks on medical image segmentation tasks typically relies on large labeled datasets for model training.
Joint Semi-supervised 3D Super-Resolution and Segmentation with Mixed Adversarial Gaussian Domain Adaptation
no code implementations • 16 Jul 2021 • Nicolo Savioli, Antonio de Marvao, Wenjia Bai, Shuo Wang, Stuart A. Cook, Calvin W. L. Chin, Daniel Rueckert, Declan P. O'Regan
Optimising the analysis of cardiac structure and function requires accurate 3D representations of shape and motion.
Joint Motion Correction and Super Resolution for Cardiac Segmentation via Latent Optimisation
no code implementations • 8 Jul 2021 • Shuo Wang, Chen Qin, Nicolo Savioli, Chen Chen, Declan O'Regan, Stuart Cook, Yike Guo, Daniel Rueckert, Wenjia Bai
In cardiac magnetic resonance (CMR) imaging, a 3D high-resolution segmentation of the heart is essential for detailed description of its anatomical structures.
Cooperative Training and Latent Space Data Augmentation for Robust Medical Image Segmentation
2 code implementations • 2 Jul 2021 • Chen Chen, Kerstin Hammernik, Cheng Ouyang, Chen Qin, Wenjia Bai, Daniel Rueckert
In this paper, we present a cooperative framework for training image segmentation models and a latent space augmentation method for generating hard examples.
A General Framework for Revealing Human Mind with auto-encoding GANs
no code implementations • 10 Feb 2021 • Pan Wang, Rui Zhou, Shuo Wang, Ling Li, Wenjia Bai, Jialu Fan, Chunlin Li, Peter Childs, Yike Guo
For this reason, we propose an end-to-end brain decoding framework which translates brain activity into an image by latent space alignment.
Suggestive Annotation of Brain Tumour Images with Gradient-guided Sampling
no code implementations • 26 Jun 2020 • Chengliang Dai, Shuo Wang, Yuanhan Mo, Kaichen Zhou, Elsa Angelini, Yike Guo, Wenjia Bai
Machine learning has been widely adopted for medical image analysis in recent years given its promising performance in image segmentation and classification tasks.
Realistic Adversarial Data Augmentation for MR Image Segmentation
1 code implementation • 23 Jun 2020 • Chen Chen, Chen Qin, Huaqi Qiu, Cheng Ouyang, Shuo Wang, Liang Chen, Giacomo Tarroni, Wenjia Bai, Daniel Rueckert
In this work, we propose an adversarial data augmentation method for training neural networks for medical image segmentation.
Deep Generative Model-based Quality Control for Cardiac MRI Segmentation
no code implementations • 23 Jun 2020 • Shuo Wang, Giacomo Tarroni, Chen Qin, Yuanhan Mo, Chengliang Dai, Chen Chen, Ben Glocker, Yike Guo, Daniel Rueckert, Wenjia Bai
Our approach provides a real-time and model-agnostic quality control for cardiac MRI segmentation, which has the potential to be integrated into clinical image analysis workflows.
Biomechanics-informed Neural Networks for Myocardial Motion Tracking in MRI
1 code implementation • 8 Jun 2020 • Chen Qin, Shuo Wang, Chen Chen, Huaqi Qiu, Wenjia Bai, Daniel Rueckert
The learnt VAE regulariser then can be coupled with any deep learning based registration network to regularise the solution space to be biomechanically plausible.
A Global Benchmark of Algorithms for Segmenting Late Gadolinium-Enhanced Cardiac Magnetic Resonance Imaging
1 code implementation • 26 Apr 2020 • Zhaohan Xiong, Qing Xia, Zhiqiang Hu, Ning Huang, Cheng Bian, Yefeng Zheng, Sulaiman Vesal, Nishant Ravikumar, Andreas Maier, Xin Yang, Pheng-Ann Heng, Dong Ni, Caizi Li, Qianqian Tong, Weixin Si, Elodie Puybareau, Younes Khoudli, Thierry Geraud, Chen Chen, Wenjia Bai, Daniel Rueckert, Lingchao Xu, Xiahai Zhuang, Xinzhe Luo, Shuman Jia, Maxime Sermesant, Yashu Liu, Kuanquan Wang, Davide Borra, Alessandro Masci, Cristiana Corsi, Coen de Vente, Mitko Veta, Rashed Karim, Chandrakanth Jayachandran Preetha, Sandy Engelhardt, Menyun Qiao, Yuanyuan Wang, Qian Tao, Marta Nunez-Garcia, Oscar Camara, Nicolo Savioli, Pablo Lamata, Jichao Zhao
Segmentation of cardiac images, particularly late gadolinium-enhanced magnetic resonance imaging (LGE-MRI) widely used for visualizing diseased cardiac structures, is a crucial first step for clinical diagnosis and treatment.
Efficient Deep Representation Learning by Adaptive Latent Space Sampling
no code implementations • 19 Mar 2020 • Yuanhan Mo, Shuo Wang, Chengliang Dai, Rui Zhou, Zhongzhao Teng, Wenjia Bai, Yike Guo
Supervised deep learning requires a large amount of training samples with annotations (e. g. label class for classification task, pixel- or voxel-wised label map for segmentation tasks), which are expensive and time-consuming to obtain.
Suggestive Labelling for Medical Image Analysis by Adaptive Latent Space Sampling
no code implementations • MIDL 2019 • Yuanhan Mo, Shuo Wang, Chengliang Dai, Zhongzhao Teng, Wenjia Bai, Yike Guo
Supervised deep learning for medical imaging analysis requires a large amount of training samples with annotations (e. g. label class for classification task, pixel- or voxel-wised label map for medical segmentation tasks), which are expensive and time-consuming to obtain.
Automatic Brain Tumour Segmentation and Biophysics-Guided Survival Prediction
no code implementations • 19 Nov 2019 • Shuo Wang, Chengliang Dai, Yuanhan Mo, Elsa Angelini, Yike Guo, Wenjia Bai
Gliomas are the most common malignant brain tumourswith intrinsic heterogeneity.
Deep learning for cardiac image segmentation: A review
no code implementations • 9 Nov 2019 • Chen Chen, Chen Qin, Huaqi Qiu, Giacomo Tarroni, Jinming Duan, Wenjia Bai, Daniel Rueckert
Deep learning has become the most widely used approach for cardiac image segmentation in recent years.
Transfer Learning from Partial Annotations for Whole Brain Segmentation
no code implementations • 28 Aug 2019 • Chengliang Dai, Yuanhan Mo, Elsa Angelini, Yike Guo, Wenjia Bai
Brain MR image segmentation is a key task in neuroimaging studies.
Unsupervised Multi-modal Style Transfer for Cardiac MR Segmentation
no code implementations • 20 Aug 2019 • Chen Chen, Cheng Ouyang, Giacomo Tarroni, Jo Schlemper, Huaqi Qiu, Wenjia Bai, Daniel Rueckert
In this work, we present a fully automatic method to segment cardiac structures from late-gadolinium enhanced (LGE) images without using labelled LGE data for training, but instead by transferring the anatomical knowledge and features learned on annotated balanced steady-state free precession (bSSFP) images, which are easier to acquire.
Joint Motion Estimation and Segmentation from Undersampled Cardiac MR Image
no code implementations • 20 Aug 2019 • Chen Qin, Wenjia Bai, Jo Schlemper, Steffen E. Petersen, Stefan K. Piechnik, Stefan Neubauer, Daniel Rueckert
Accelerating the acquisition of magnetic resonance imaging (MRI) is a challenging problem, and many works have been proposed to reconstruct images from undersampled k-space data.
Learning Shape Priors for Robust Cardiac MR Segmentation from Multi-view Images
no code implementations • 23 Jul 2019 • Chen Chen, Carlo Biffi, Giacomo Tarroni, Steffen Petersen, Wenjia Bai, Daniel Rueckert
Cardiac MR image segmentation is essential for the morphological and functional analysis of the heart.
VS-Net: Variable splitting network for accelerated parallel MRI reconstruction
1 code implementation • 19 Jul 2019 • Jinming Duan, Jo Schlemper, Chen Qin, Cheng Ouyang, Wenjia Bai, Carlo Biffi, Ghalib Bello, Ben Statton, Declan P. O'Regan, Daniel Rueckert
In this work, we propose a deep learning approach for parallel magnetic resonance imaging (MRI) reconstruction, termed a variable splitting network (VS-Net), for an efficient, high-quality reconstruction of undersampled multi-coil MR data.
Self-Supervised Learning for Cardiac MR Image Segmentation by Anatomical Position Prediction
no code implementations • 5 Jul 2019 • Wenjia Bai, Chen Chen, Giacomo Tarroni, Jinming Duan, Florian Guitton, Steffen E. Petersen, Yike Guo, Paul M. Matthews, Daniel Rueckert
In the recent years, convolutional neural networks have transformed the field of medical image analysis due to their capacity to learn discriminative image features for a variety of classification and regression tasks.
Improving the generalizability of convolutional neural network-based segmentation on CMR images
1 code implementation • 2 Jul 2019 • Chen Chen, Wenjia Bai, Rhodri H. Davies, Anish N. Bhuva, Charlotte Manisty, James C. Moon, Nay Aung, Aaron M. Lee, Mihir M. Sanghvi, Kenneth Fung, Jose Miguel Paiva, Steffen E. Petersen, Elena Lukaschuk, Stefan K. Piechnik, Stefan Neubauer, Daniel Rueckert
We demonstrate that a neural network trained on a single-site single-scanner dataset from the UK Biobank can be successfully applied to segmenting cardiac MR images across different sites and different scanners without substantial loss of accuracy.
Explainable Anatomical Shape Analysis through Deep Hierarchical Generative Models
1 code implementation • 28 Jun 2019 • Carlo Biffi, Juan J. Cerrolaza, Giacomo Tarroni, Wenjia Bai, Antonio de Marvao, Ozan Oktay, Christian Ledig, Loic Le Folgoc, Konstantinos Kamnitsas, Georgia Doumou, Jinming Duan, Sanjay K. Prasad, Stuart A. Cook, Declan P. O'Regan, Daniel Rueckert
At the highest level of this hierarchy, a two-dimensional latent space is simultaneously optimised to discriminate distinct clinical conditions, enabling the direct visualisation of the classification space.
Automated Quality Control in Image Segmentation: Application to the UK Biobank Cardiac MR Imaging Study
no code implementations • 27 Jan 2019 • Robert Robinson, Vanya V. Valindria, Wenjia Bai, Ozan Oktay, Bernhard Kainz, Hideaki Suzuki, Mihir M. Sanghvi, Nay Aung, Jos$é$ Miguel Paiva, Filip Zemrak, Kenneth Fung, Elena Lukaschuk, Aaron M. Lee, Valentina Carapella, Young Jin Kim, Stefan K. Piechnik, Stefan Neubauer, Steffen E. Petersen, Chris Page, Paul M. Matthews, Daniel Rueckert, Ben Glocker
Methods: To overcome this challenge, we explore an approach for predicting segmentation quality based on Reverse Classification Accuracy, which enables us to discriminate between successful and failed segmentations on a per-cases basis.
Multi-Task Learning for Left Atrial Segmentation on GE-MRI
1 code implementation • 31 Oct 2018 • Chen Chen, Wenjia Bai, Daniel Rueckert
Segmentation of the left atrium (LA) is crucial for assessing its anatomy in both pre-operative atrial fibrillation (AF) ablation planning and post-operative follow-up studies.
A Comprehensive Approach for Learning-based Fully-Automated Inter-slice Motion Correction for Short-Axis Cine Cardiac MR Image Stacks
no code implementations • 3 Oct 2018 • Giacomo Tarroni, Ozan Oktay, Matthew Sinclair, Wenjia Bai, Andreas Schuh, Hideaki Suzuki, Antonio de Marvao, Declan O'Regan, Stuart Cook, Daniel Rueckert
If long axis (LA) images are available, PSMs are generated for them and combined to create the target PSM; if not, the target PSM is produced from the same stack using a 3D model trained from motion-free stacks.
Automatic 3D bi-ventricular segmentation of cardiac images by a shape-refined multi-task deep learning approach
1 code implementation • 26 Aug 2018 • Jinming Duan, Ghalib Bello, Jo Schlemper, Wenjia Bai, Timothy J. W. Dawes, Carlo Biffi, Antonio de Marvao, Georgia Doumou, Declan P. O'Regan, Daniel Rueckert
The proposed pipeline is fully automated, due to network's ability to infer landmarks, which are then used downstream in the pipeline to initialise atlas propagation.
Recurrent neural networks for aortic image sequence segmentation with sparse annotations
no code implementations • 1 Aug 2018 • Wenjia Bai, Hideaki Suzuki, Chen Qin, Giacomo Tarroni, Ozan Oktay, Paul M. Matthews, Daniel Rueckert
In this work, we propose an image sequence segmentation algorithm by combining a fully convolutional network with a recurrent neural network, which incorporates both spatial and temporal information into the segmentation task.
Deep nested level sets: Fully automated segmentation of cardiac MR images in patients with pulmonary hypertension
no code implementations • 27 Jul 2018 • Jinming Duan, Jo Schlemper, Wenjia Bai, Timothy J. W. Dawes, Ghalib Bello, Georgia Doumou, Antonio de Marvao, Declan P. O'Regan, Daniel Rueckert
In this paper we introduce a novel and accurate optimisation method for segmentation of cardiac MR (CMR) images in patients with pulmonary hypertension (PH).
Learning Interpretable Anatomical Features Through Deep Generative Models: Application to Cardiac Remodeling
1 code implementation • 18 Jul 2018 • Carlo Biffi, Ozan Oktay, Giacomo Tarroni, Wenjia Bai, Antonio de Marvao, Georgia Doumou, Martin Rajchl, Reem Bedair, Sanjay Prasad, Stuart Cook, Declan O'Regan, Daniel Rueckert
However, current approaches to the diagnosis of cardiovascular diseases often rely on subjective human assessment as well as manual analysis of medical images.
Real-time Prediction of Segmentation Quality
no code implementations • 16 Jun 2018 • Robert Robinson, Ozan Oktay, Wenjia Bai, Vanya Valindria, Mihir Sanghvi, Nay Aung, José Paiva, Filip Zemrak, Kenneth Fung, Elena Lukaschuk, Aaron Lee, Valentina Carapella, Young Jin Kim, Bernhard Kainz, Stefan Piechnik, Stefan Neubauer, Steffen Petersen, Chris Page, Daniel Rueckert, Ben Glocker
Recent advances in deep learning based image segmentation methods have enabled real-time performance with human-level accuracy.
Joint Learning of Motion Estimation and Segmentation for Cardiac MR Image Sequences
1 code implementation • 11 Jun 2018 • Chen Qin, Wenjia Bai, Jo Schlemper, Steffen E. Petersen, Stefan K. Piechnik, Stefan Neubauer, Daniel Rueckert
Cardiac motion estimation and segmentation play important roles in quantitatively assessing cardiac function and diagnosing cardiovascular diseases.
Automatic View Planning with Multi-scale Deep Reinforcement Learning Agents
no code implementations • 8 Jun 2018 • Amir Alansary, Loic Le Folgoc, Ghislain Vaillant, Ozan Oktay, Yuanwei Li, Wenjia Bai, Jonathan Passerat-Palmbach, Ricardo Guerrero, Konstantinos Kamnitsas, Benjamin Hou, Steven McDonagh, Ben Glocker, Bernhard Kainz, Daniel Rueckert
Navigating through target anatomy to find the required view plane is tedious and operator-dependent.
Domain Adaptation for MRI Organ Segmentation using Reverse Classification Accuracy
1 code implementation • 1 Jun 2018 • Vanya V. Valindria, Ioannis Lavdas, Wenjia Bai, Konstantinos Kamnitsas, Eric O. Aboagye, Andrea G. Rockall, Daniel Rueckert, Ben Glocker
The variations in multi-center data in medical imaging studies have brought the necessity of domain adaptation.
Human-level Performance On Automatic Head Biometrics In Fetal Ultrasound Using Fully Convolutional Neural Networks
no code implementations • 24 Apr 2018 • Matthew Sinclair, Christian F. Baumgartner, Jacqueline Matthew, Wenjia Bai, Juan Cerrolaza Martinez, Yuanwei Li, Sandra Smith, Caroline L. Knight, Bernhard Kainz, Jo Hajnal, Andrew P. King, Daniel Rueckert
Measurement of head biometrics from fetal ultrasonography images is of key importance in monitoring the healthy development of fetuses.
Learning-Based Quality Control for Cardiac MR Images
no code implementations • 25 Mar 2018 • Giacomo Tarroni, Ozan Oktay, Wenjia Bai, Andreas Schuh, Hideaki Suzuki, Jonathan Passerat-Palmbach, Antonio de Marvao, Declan P. O'Regan, Stuart Cook, Ben Glocker, Paul M. Matthews, Daniel Rueckert
The results show the capability of the proposed pipeline to correctly detect incomplete or corrupted scans (e. g. on UK Biobank, sensitivity and specificity respectively 88% and 99% for heart coverage estimation, 85% and 95% for motion detection), allowing their exclusion from the analysed dataset or the triggering of a new acquisition.
Ensembles of Multiple Models and Architectures for Robust Brain Tumour Segmentation
no code implementations • 4 Nov 2017 • Konstantinos Kamnitsas, Wenjia Bai, Enzo Ferrante, Steven McDonagh, Matthew Sinclair, Nick Pawlowski, Martin Rajchl, Matthew Lee, Bernhard Kainz, Daniel Rueckert, Ben Glocker
Deep learning approaches such as convolutional neural nets have consistently outperformed previous methods on challenging tasks such as dense, semantic segmentation.
Automated cardiovascular magnetic resonance image analysis with fully convolutional networks
1 code implementation • 25 Oct 2017 • Wenjia Bai, Matthew Sinclair, Giacomo Tarroni, Ozan Oktay, Martin Rajchl, Ghislain Vaillant, Aaron M. Lee, Nay Aung, Elena Lukaschuk, Mihir M. Sanghvi, Filip Zemrak, Kenneth Fung, Jose Miguel Paiva, Valentina Carapella, Young Jin Kim, Hideaki Suzuki, Bernhard Kainz, Paul M. Matthews, Steffen E. Petersen, Stefan K. Piechnik, Stefan Neubauer, Ben Glocker, Daniel Rueckert
By combining FCN with a large-scale annotated dataset, the proposed automated method achieves a high performance on par with human experts in segmenting the LV and RV on short-axis CMR images and the left atrium (LA) and right atrium (RA) on long-axis CMR images.
Anatomically Constrained Neural Networks (ACNN): Application to Cardiac Image Enhancement and Segmentation
1 code implementation • 22 May 2017 • Ozan Oktay, Enzo Ferrante, Konstantinos Kamnitsas, Mattias Heinrich, Wenjia Bai, Jose Caballero, Stuart Cook, Antonio de Marvao, Timothy Dawes, Declan O'Regan, Bernhard Kainz, Ben Glocker, Daniel Rueckert
However, in most recent and promising techniques such as CNN based segmentation it is not obvious how to incorporate such prior knowledge.
Reverse Classification Accuracy: Predicting Segmentation Performance in the Absence of Ground Truth
no code implementations • 11 Feb 2017 • Vanya V. Valindria, Ioannis Lavdas, Wenjia Bai, Konstantinos Kamnitsas, Eric O. Aboagye, Andrea G. Rockall, Daniel Rueckert, Ben Glocker
In RCA we take the predicted segmentation from a new image to train a reverse classifier which is evaluated on a set of reference images with available ground truth.
DeepCut: Object Segmentation from Bounding Box Annotations using Convolutional Neural Networks
no code implementations • 25 May 2016 • Martin Rajchl, Matthew C. H. Lee, Ozan Oktay, Konstantinos Kamnitsas, Jonathan Passerat-Palmbach, Wenjia Bai, Mellisa Damodaram, Mary A. Rutherford, Joseph V. Hajnal, Bernhard Kainz, Daniel Rueckert
In this paper, we propose DeepCut, a method to obtain pixelwise object segmentations given an image dataset labelled with bounding box annotations.
Multi-Atlas Segmentation using Partially Annotated Data: Methods and Annotation Strategies
no code implementations • 29 Apr 2016 • Lisa M. Koch, Martin Rajchl, Wenjia Bai, Christian F. Baumgartner, Tong Tong, Jonathan Passerat-Palmbach, Paul Aljabar, Daniel Rueckert
Multi-atlas segmentation is a widely used tool in medical image analysis, providing robust and accurate results by learning from annotated atlas datasets.
Patch-based Evaluation of Image Segmentation
no code implementations • CVPR 2014 • Christian Ledig, Wenzhe Shi, Wenjia Bai, Daniel Rueckert
The ideal similarity measure should be unbiased to segmentations of different volume and complexity, and be able to quantify and visualise segmentation bias.