Search Results for author: Ben Glocker
Found 124 papers, 71 papers with code
The Importance of Model Inspection for Better Understanding Performance Characteristics of Graph Neural Networks
1 code implementation • 2 May 2024 • Nairouz Shehata, Carolina Piçarra, Anees Kazi, Ben Glocker
This study highlights the importance of conducting comprehensive model inspection as part of comparative performance analyses.
Mitigating attribute amplification in counterfactual image generation
no code implementations • 14 Mar 2024 • Tian Xia, Mélanie Roschewitz, Fabio De Sousa Ribeiro, Charles Jones, Ben Glocker
Causal generative modelling is gaining interest in medical imaging due to its ability to answer interventional and counterfactual queries.
Counterfactual contrastive learning: robust representations via causal image synthesis
1 code implementation • 14 Mar 2024 • Melanie Roschewitz, Fabio De Sousa Ribeiro, Tian Xia, Galvin Khara, Ben Glocker
Contrastive pretraining is well-known to improve downstream task performance and model generalisation, especially in limited label settings.
Average Calibration Error: A Differentiable Loss for Improved Reliability in Image Segmentation
1 code implementation • 11 Mar 2024 • Theodore Barfoot, Luis Garcia-Peraza-Herrera, Ben Glocker, Tom Vercauteren
Using mL1-ACE, we reduce average and maximum calibration error by 45% and 55% respectively, maintaining a Dice score of 87% on the BraTS 2021 dataset.
Demystifying Variational Diffusion Models
no code implementations • 11 Jan 2024 • Fabio De Sousa Ribeiro, Ben Glocker
Despite the growing popularity of diffusion models, gaining a deep understanding of the model class remains somewhat elusive for the uninitiated in non-equilibrium statistical physics.
Robust semi-supervised segmentation with timestep ensembling diffusion models
no code implementations • 13 Nov 2023 • Margherita Rosnati, Melanie Roschewitz, Ben Glocker
Medical image segmentation is a challenging task, made more difficult by many datasets' limited size and annotations.
Analysing race and sex bias in brain age prediction
no code implementations • 19 Sep 2023 • Carolina Piçarra, Ben Glocker
With the objective of comparing the performance between subgroups, measured by the absolute prediction error, we use a Kruskal-Wallis test followed by two post-hoc Conover-Iman tests to inspect bias across race and biological sex.
Robustness Stress Testing in Medical Image Classification
1 code implementation • 14 Aug 2023 • Mobarakol Islam, Zeju Li, Ben Glocker
We conclude that progressive stress testing is a viable and important tool and should become standard practice in the clinical validation of image-based disease detection models.
Distance Matters For Improving Performance Estimation Under Covariate Shift
1 code implementation • 14 Aug 2023 • Mélanie Roschewitz, Ben Glocker
In this work, we show that taking into account distances of test samples to their expected training distribution can significantly improve performance estimation under covariate shift.
FUTURE-AI: International consensus guideline for trustworthy and deployable artificial intelligence in healthcare
no code implementations • 11 Aug 2023 • Karim Lekadir, Aasa Feragen, Abdul Joseph Fofanah, Alejandro F Frangi, Alena Buyx, Anais Emelie, Andrea Lara, Antonio R Porras, An-Wen Chan, Arcadi Navarro, Ben Glocker, Benard O Botwe, Bishesh Khanal, Brigit Beger, Carol C Wu, Celia Cintas, Curtis P Langlotz, Daniel Rueckert, Deogratias Mzurikwao, Dimitrios I Fotiadis, Doszhan Zhussupov, Enzo Ferrante, Erik Meijering, Eva Weicken, Fabio A González, Folkert W Asselbergs, Fred Prior, Gabriel P Krestin, Gary Collins, Geletaw S Tegenaw, Georgios Kaissis, Gianluca Misuraca, Gianna Tsakou, Girish Dwivedi, Haridimos Kondylakis, Harsha Jayakody, Henry C Woodruf, Hugo JWL Aerts, Ian Walsh, Ioanna Chouvarda, Irène Buvat, Islem Rekik, James Duncan, Jayashree Kalpathy-Cramer, Jihad Zahir, Jinah Park, John Mongan, Judy W Gichoya, Julia A Schnabel, Kaisar Kushibar, Katrine Riklund, Kensaku MORI, Kostas Marias, Lameck M Amugongo, Lauren A Fromont, Lena Maier-Hein, Leonor Cerdá Alberich, Leticia Rittner, Lighton Phiri, Linda Marrakchi-Kacem, Lluís Donoso-Bach, Luis Martí-Bonmatí, M Jorge Cardoso, Maciej Bobowicz, Mahsa Shabani, Manolis Tsiknakis, Maria A Zuluaga, Maria Bielikova, Marie-Christine Fritzsche, Marius George Linguraru, Markus Wenzel, Marleen de Bruijne, Martin G Tolsgaard, Marzyeh Ghassemi, Md Ashrafuzzaman, Melanie Goisauf, Mohammad Yaqub, Mohammed Ammar, Mónica Cano Abadía, Mukhtar M E Mahmoud, Mustafa Elattar, Nicola Rieke, Nikolaos Papanikolaou, Noussair Lazrak, Oliver Díaz, Olivier Salvado, Oriol Pujol, Ousmane Sall, Pamela Guevara, Peter Gordebeke, Philippe Lambin, Pieta Brown, Purang Abolmaesumi, Qi Dou, Qinghua Lu, Richard Osuala, Rose Nakasi, S Kevin Zhou, Sandy Napel, Sara Colantonio, Shadi Albarqouni, Smriti Joshi, Stacy Carter, Stefan Klein, Steffen E Petersen, Susanna Aussó, Suyash Awate, Tammy Riklin Raviv, Tessa Cook, Tinashe E M Mutsvangwa, Wendy A Rogers, Wiro J Niessen, Xènia Puig-Bosch, Yi Zeng, Yunusa G Mohammed, Yves Saint James Aquino, Zohaib Salahuddin, Martijn P A Starmans
This work describes the FUTURE-AI guideline as the first international consensus framework for guiding the development and deployment of trustworthy AI tools in healthcare.
No Fair Lunch: A Causal Perspective on Dataset Bias in Machine Learning for Medical Imaging
no code implementations • 31 Jul 2023 • Charles Jones, Daniel C. Castro, Fabio De Sousa Ribeiro, Ozan Oktay, Melissa McCradden, Ben Glocker
As machine learning methods gain prominence within clinical decision-making, addressing fairness concerns becomes increasingly urgent.
Grounded Object Centric Learning
no code implementations • 18 Jul 2023 • Avinash Kori, Francesco Locatello, Fabio De Sousa Ribeiro, Francesca Toni, Ben Glocker
The extraction of modular object-centric representations for downstream tasks is an emerging area of research.
A Causal Ordering Prior for Unsupervised Representation Learning
no code implementations • 11 Jul 2023 • Avinash Kori, Pedro Sanchez, Konstantinos Vilouras, Ben Glocker, Sotirios A. Tsaftaris
Unsupervised representation learning with variational inference relies heavily on independence assumptions over latent variables.
The Role of Subgroup Separability in Group-Fair Medical Image Classification
1 code implementation • 6 Jul 2023 • Charles Jones, Mélanie Roschewitz, Ben Glocker
We investigate performance disparities in deep classifiers.
High Fidelity Image Counterfactuals with Probabilistic Causal Models
1 code implementation • 27 Jun 2023 • Fabio De Sousa Ribeiro, Tian Xia, Miguel Monteiro, Nick Pawlowski, Ben Glocker
We present a general causal generative modelling framework for accurate estimation of high fidelity image counterfactuals with deep structural causal models.
Joint Optimization of Class-Specific Training- and Test-Time Data Augmentation in Segmentation
1 code implementation • 30 May 2023 • Zeju Li, Konstantinos Kamnitsas, Qi Dou, Chen Qin, Ben Glocker
We demonstrate the effectiveness of our method on four medical image segmentation tasks across different scenarios with two state-of-the-art segmentation models, DeepMedic and nnU-Net.
Measuring axiomatic soundness of counterfactual image models
1 code implementation • 2 Mar 2023 • Miguel Monteiro, Fabio De Sousa Ribeiro, Nick Pawlowski, Daniel C. Castro, Ben Glocker
We present a general framework for evaluating image counterfactuals.
Understanding metric-related pitfalls in image analysis validation
no code implementations • 3 Feb 2023 • Annika Reinke, Minu D. Tizabi, Michael Baumgartner, Matthias Eisenmann, Doreen Heckmann-Nötzel, A. Emre Kavur, Tim Rädsch, Carole H. Sudre, Laura Acion, Michela Antonelli, Tal Arbel, Spyridon Bakas, Arriel Benis, Matthew Blaschko, Florian Buettner, M. Jorge Cardoso, Veronika Cheplygina, Jianxu Chen, Evangelia Christodoulou, Beth A. Cimini, Gary S. Collins, Keyvan Farahani, Luciana Ferrer, Adrian Galdran, Bram van Ginneken, Ben Glocker, Patrick Godau, Robert Haase, Daniel A. Hashimoto, Michael M. Hoffman, Merel Huisman, Fabian Isensee, Pierre Jannin, Charles E. Kahn, Dagmar Kainmueller, Bernhard Kainz, Alexandros Karargyris, Alan Karthikesalingam, Hannes Kenngott, Jens Kleesiek, Florian Kofler, Thijs Kooi, Annette Kopp-Schneider, Michal Kozubek, Anna Kreshuk, Tahsin Kurc, Bennett A. Landman, Geert Litjens, Amin Madani, Klaus Maier-Hein, Anne L. Martel, Peter Mattson, Erik Meijering, Bjoern Menze, Karel G. M. Moons, Henning Müller, Brennan Nichyporuk, Felix Nickel, Jens Petersen, Susanne M. Rafelski, Nasir Rajpoot, Mauricio Reyes, Michael A. Riegler, Nicola Rieke, Julio Saez-Rodriguez, Clara I. Sánchez, Shravya Shetty, Maarten van Smeden, Ronald M. Summers, Abdel A. Taha, Aleksei Tiulpin, Sotirios A. Tsaftaris, Ben van Calster, Gaël Varoquaux, Manuel Wiesenfarth, Ziv R. Yaniv, Paul F. Jäger, Lena Maier-Hein
Validation metrics are key for the reliable tracking of scientific progress and for bridging the current chasm between artificial intelligence (AI) research and its translation into practice.
Paced-Curriculum Distillation with Prediction and Label Uncertainty for Image Segmentation
1 code implementation • 2 Feb 2023 • Mobarakol Islam, Lalithkumar Seenivasan, S. P. Sharan, V. K. Viekash, Bhavesh Gupta, Ben Glocker, Hongliang Ren
Purpose: In curriculum learning, the idea is to train on easier samples first and gradually increase the difficulty, while in self-paced learning, a pacing function defines the speed to adapt the training progress.
Image To Tree with Recursive Prompting
no code implementations • 1 Jan 2023 • James Batten, Matthew Sinclair, Ben Glocker, Michiel Schaap
Extracting complex structures from grid-based data is a common key step in automated medical image analysis.
Confidence-Aware Paced-Curriculum Learning by Label Smoothing for Surgical Scene Understanding
1 code implementation • 22 Dec 2022 • Mengya Xu, Mobarakol Islam, Ben Glocker, Hongliang Ren
In this work, we design a paced curriculum by label smoothing (P-CBLS) using paced learning with uniform label smoothing (ULS) for classification tasks and fuse uniform and spatially varying label smoothing (SVLS) for semantic segmentation tasks in a curriculum manner.
Context Label Learning: Improving Background Class Representations in Semantic Segmentation
1 code implementation • 16 Dec 2022 • Zeju Li, Konstantinos Kamnitsas, Cheng Ouyang, Chen Chen, Ben Glocker
The results demonstrate that CoLab can guide the segmentation model to map the logits of background samples away from the decision boundary, resulting in significantly improved segmentation accuracy.
Analysing the effectiveness of a generative model for semi-supervised medical image segmentation
no code implementations • 3 Nov 2022 • Margherita Rosnati, Fabio De Sousa Ribeiro, Miguel Monteiro, Daniel Coelho de Castro, Ben Glocker
In such settings, semi-supervised learning (SSL) attempts to leverage the abundance of unlabelled data to obtain more robust and reliable models.
A Comparative Study of Graph Neural Networks for Shape Classification in Neuroimaging
1 code implementation • 29 Oct 2022 • Nairouz Shehata, Wulfie Bain, Ben Glocker
Graph neural networks have emerged as a promising approach for the analysis of non-Euclidean data such as meshes.
Explaining Image Classification with Visual Debates
1 code implementation • 17 Oct 2022 • Avinash Kori, Ben Glocker, Francesca Toni
An effective way to obtain different perspectives on any given topic is by conducting a debate, where participants argue for and against the topic.
Frequency Dropout: Feature-Level Regularization via Randomized Filtering
no code implementations • 20 Sep 2022 • Mobarakol Islam, Ben Glocker
Both high and low frequencies can be characteristic of the underlying noise distribution caused by the image acquisition rather than in relation to the task-relevant information about the image content.
Risk of Bias in Chest Radiography Deep Learning Foundation Models
2 code implementations • 7 Sep 2022 • Ben Glocker, Charles Jones, Melanie Roschewitz, Stefan Winzeck
Purpose: To analyze a recently published chest radiography foundation model for the presence of biases that could lead to subgroup performance disparities across biological sex and race.
Deep Structural Causal Shape Models
no code implementations • 23 Aug 2022 • Rajat Rasal, Daniel C. Castro, Nick Pawlowski, Ben Glocker
Causal reasoning provides a language to ask important interventional and counterfactual questions beyond purely statistical association.
Evaluation of 3D GANs for Lung Tissue Modelling in Pulmonary CT
1 code implementation • 17 Aug 2022 • Sam Ellis, Octavio E. Martinez Manzanera, Vasileios Baltatzis, Ibrahim Nawaz, Arjun Nair, Loïc le Folgoc, Sujal Desai, Ben Glocker, Julia A. Schnabel
This makes high-quality GANs useful for unsupervised anomaly detection in medical imaging.
Automatic lesion analysis for increased efficiency in outcome prediction of traumatic brain injury
no code implementations • 8 Aug 2022 • Margherita Rosnati, Eyal Soreq, Miguel Monteiro, Lucia Li, Neil S. N. Graham, Karl Zimmerman, Carlotta Rossi, Greta Carrara, Guido Bertolini, David J. Sharp, Ben Glocker
We compare the predictive power of our proposed features to the Marshall score, independently and when paired with classic TBI biomarkers.
Estimating Model Performance under Domain Shifts with Class-Specific Confidence Scores
1 code implementation • 20 Jul 2022 • Zeju Li, Konstantinos Kamnitsas, Mobarakol Islam, Chen Chen, Ben Glocker
If we could estimate the performance that a pre-trained model would achieve on data from a specific deployment setting, for example a certain clinic, we could judge whether the model could safely be deployed or if its performance degrades unacceptably on the specific data.
GLANCE: Global to Local Architecture-Neutral Concept-based Explanations
no code implementations • 5 Jul 2022 • Avinash Kori, Ben Glocker, Francesca Toni
Specifically, we provide a generator to visualize the `effect' of interactions among features in latent space and draw feature importance therefrom as local explanations.
Vector Quantisation for Robust Segmentation
1 code implementation • 5 Jul 2022 • Ainkaran Santhirasekaram, Avinash Kori, Mathias Winkler, Andrea Rockall, Ben Glocker
The reliability of segmentation models in the medical domain depends on the model's robustness to perturbations in the input space.
Hierarchical Symbolic Reasoning in Hyperbolic Space for Deep Discriminative Models
1 code implementation • 5 Jul 2022 • Ainkaran Santhirasekaram, Avinash Kori, Andrea Rockall, Mathias Winkler, Francesca Toni, Ben Glocker
We achieve this by using the natural properties of \emph{hyperbolic geometry} to more efficiently model a hierarchy of symbolic features and generate \emph{hierarchical symbolic rules} as part of our explanations.
Distributional Gaussian Processes Layers for Out-of-Distribution Detection
no code implementations • 27 Jun 2022 • Sebastian G. Popescu, David J. Sharp, James H. Cole, Konstantinos Kamnitsas, Ben Glocker
Moreover, by applying the same segmentation model to out-of-distribution data (i. e., images with pathology such as brain tumors), we show that our uncertainty estimates result in out-of-distribution detection that outperforms the capabilities of previous Bayesian networks and reconstruction-based approaches that learn normative distributions.
Metrics reloaded: Recommendations for image analysis validation
1 code implementation • 3 Jun 2022 • Lena Maier-Hein, Annika Reinke, Patrick Godau, Minu D. Tizabi, Florian Buettner, Evangelia Christodoulou, Ben Glocker, Fabian Isensee, Jens Kleesiek, Michal Kozubek, Mauricio Reyes, Michael A. Riegler, Manuel Wiesenfarth, A. Emre Kavur, Carole H. Sudre, Michael Baumgartner, Matthias Eisenmann, Doreen Heckmann-Nötzel, Tim Rädsch, Laura Acion, Michela Antonelli, Tal Arbel, Spyridon Bakas, Arriel Benis, Matthew Blaschko, M. Jorge Cardoso, Veronika Cheplygina, Beth A. Cimini, Gary S. Collins, Keyvan Farahani, Luciana Ferrer, Adrian Galdran, Bram van Ginneken, Robert Haase, Daniel A. Hashimoto, Michael M. Hoffman, Merel Huisman, Pierre Jannin, Charles E. Kahn, Dagmar Kainmueller, Bernhard Kainz, Alexandros Karargyris, Alan Karthikesalingam, Hannes Kenngott, Florian Kofler, Annette Kopp-Schneider, Anna Kreshuk, Tahsin Kurc, Bennett A. Landman, Geert Litjens, Amin Madani, Klaus Maier-Hein, Anne L. Martel, Peter Mattson, Erik Meijering, Bjoern Menze, Karel G. M. Moons, Henning Müller, Brennan Nichyporuk, Felix Nickel, Jens Petersen, Nasir Rajpoot, Nicola Rieke, Julio Saez-Rodriguez, Clara I. Sánchez, Shravya Shetty, Maarten van Smeden, Ronald M. Summers, Abdel A. Taha, Aleksei Tiulpin, Sotirios A. Tsaftaris, Ben van Calster, Gaël Varoquaux, Paul F. Jäger
The framework was developed in a multi-stage Delphi process and is based on the novel concept of a problem fingerprint - a structured representation of the given problem that captures all aspects that are relevant for metric selection, from the domain interest to the properties of the target structure(s), data set and algorithm output.
Failure Detection in Medical Image Classification: A Reality Check and Benchmarking Testbed
1 code implementation • 27 May 2022 • Melanie Bernhardt, Fabio De Sousa Ribeiro, Ben Glocker
Failure detection in automated image classification is a critical safeguard for clinical deployment.
Structured Uncertainty in the Observation Space of Variational Autoencoders
1 code implementation • 25 May 2022 • James Langley, Miguel Monteiro, Charles Jones, Nick Pawlowski, Ben Glocker
In contrast, improving the model for the observational distribution is rarely considered and typically defaults to a pixel-wise independent categorical or normal distribution.
Federated Learning Enables Big Data for Rare Cancer Boundary Detection
1 code implementation • 22 Apr 2022 • Sarthak Pati, Ujjwal Baid, Brandon Edwards, Micah Sheller, Shih-han Wang, G Anthony Reina, Patrick Foley, Alexey Gruzdev, Deepthi Karkada, Christos Davatzikos, Chiharu Sako, Satyam Ghodasara, Michel Bilello, Suyash Mohan, Philipp Vollmuth, Gianluca Brugnara, Chandrakanth J Preetha, Felix Sahm, Klaus Maier-Hein, Maximilian Zenk, Martin Bendszus, Wolfgang Wick, Evan Calabrese, Jeffrey Rudie, Javier Villanueva-Meyer, Soonmee Cha, Madhura Ingalhalikar, Manali Jadhav, Umang Pandey, Jitender Saini, John Garrett, Matthew Larson, Robert Jeraj, Stuart Currie, Russell Frood, Kavi Fatania, Raymond Y Huang, Ken Chang, Carmen Balana, Jaume Capellades, Josep Puig, Johannes Trenkler, Josef Pichler, Georg Necker, Andreas Haunschmidt, Stephan Meckel, Gaurav Shukla, Spencer Liem, Gregory S Alexander, Joseph Lombardo, Joshua D Palmer, Adam E Flanders, Adam P Dicker, Haris I Sair, Craig K Jones, Archana Venkataraman, Meirui Jiang, Tiffany Y So, Cheng Chen, Pheng Ann Heng, Qi Dou, Michal Kozubek, Filip Lux, Jan Michálek, Petr Matula, Miloš Keřkovský, Tereza Kopřivová, Marek Dostál, Václav Vybíhal, Michael A Vogelbaum, J Ross Mitchell, Joaquim Farinhas, Joseph A Maldjian, Chandan Ganesh Bangalore Yogananda, Marco C Pinho, Divya Reddy, James Holcomb, Benjamin C Wagner, Benjamin M Ellingson, Timothy F Cloughesy, Catalina Raymond, Talia Oughourlian, Akifumi Hagiwara, Chencai Wang, Minh-Son To, Sargam Bhardwaj, Chee Chong, Marc Agzarian, Alexandre Xavier Falcão, Samuel B Martins, Bernardo C A Teixeira, Flávia Sprenger, David Menotti, Diego R Lucio, Pamela Lamontagne, Daniel Marcus, Benedikt Wiestler, Florian Kofler, Ivan Ezhov, Marie Metz, Rajan Jain, Matthew Lee, Yvonne W Lui, Richard McKinley, Johannes Slotboom, Piotr Radojewski, Raphael Meier, Roland Wiest, Derrick Murcia, Eric Fu, Rourke Haas, John Thompson, David Ryan Ormond, Chaitra Badve, Andrew E Sloan, Vachan Vadmal, Kristin Waite, Rivka R Colen, Linmin Pei, Murat AK, Ashok Srinivasan, J Rajiv Bapuraj, Arvind Rao, Nicholas Wang, Ota Yoshiaki, Toshio Moritani, Sevcan Turk, Joonsang Lee, Snehal Prabhudesai, Fanny Morón, Jacob Mandel, Konstantinos Kamnitsas, Ben Glocker, Luke V M Dixon, Matthew Williams, Peter Zampakis, Vasileios Panagiotopoulos, Panagiotis Tsiganos, Sotiris Alexiou, Ilias Haliassos, Evangelia I Zacharaki, Konstantinos Moustakas, Christina Kalogeropoulou, Dimitrios M Kardamakis, Yoon Seong Choi, Seung-Koo Lee, Jong Hee Chang, Sung Soo Ahn, Bing Luo, Laila Poisson, Ning Wen, Pallavi Tiwari, Ruchika Verma, Rohan Bareja, Ipsa Yadav, Jonathan Chen, Neeraj Kumar, Marion Smits, Sebastian R van der Voort, Ahmed Alafandi, Fatih Incekara, Maarten MJ Wijnenga, Georgios Kapsas, Renske Gahrmann, Joost W Schouten, Hendrikus J Dubbink, Arnaud JPE Vincent, Martin J van den Bent, Pim J French, Stefan Klein, Yading Yuan, Sonam Sharma, Tzu-Chi Tseng, Saba Adabi, Simone P Niclou, Olivier Keunen, Ann-Christin Hau, Martin Vallières, David Fortin, Martin Lepage, Bennett Landman, Karthik Ramadass, Kaiwen Xu, Silky Chotai, Lola B Chambless, Akshitkumar Mistry, Reid C Thompson, Yuriy Gusev, Krithika Bhuvaneshwar, Anousheh Sayah, Camelia Bencheqroun, Anas Belouali, Subha Madhavan, Thomas C Booth, Alysha Chelliah, Marc Modat, Haris Shuaib, Carmen Dragos, Aly Abayazeed, Kenneth Kolodziej, Michael Hill, Ahmed Abbassy, Shady Gamal, Mahmoud Mekhaimar, Mohamed Qayati, Mauricio Reyes, Ji Eun Park, Jihye Yun, Ho Sung Kim, Abhishek Mahajan, Mark Muzi, Sean Benson, Regina G H Beets-Tan, Jonas Teuwen, Alejandro Herrera-Trujillo, Maria Trujillo, William Escobar, Ana Abello, Jose Bernal, Jhon Gómez, Joseph Choi, Stephen Baek, Yusung Kim, Heba Ismael, Bryan Allen, John M Buatti, Aikaterini Kotrotsou, Hongwei Li, Tobias Weiss, Michael Weller, Andrea Bink, Bertrand Pouymayou, Hassan F Shaykh, Joel Saltz, Prateek Prasanna, Sampurna Shrestha, Kartik M Mani, David Payne, Tahsin Kurc, Enrique Pelaez, Heydy Franco-Maldonado, Francis Loayza, Sebastian Quevedo, Pamela Guevara, Esteban Torche, Cristobal Mendoza, Franco Vera, Elvis Ríos, Eduardo López, Sergio A Velastin, Godwin Ogbole, Dotun Oyekunle, Olubunmi Odafe-Oyibotha, Babatunde Osobu, Mustapha Shu'aibu, Adeleye Dorcas, Mayowa Soneye, Farouk Dako, Amber L Simpson, Mohammad Hamghalam, Jacob J Peoples, Ricky Hu, Anh Tran, Danielle Cutler, Fabio Y Moraes, Michael A Boss, James Gimpel, Deepak Kattil Veettil, Kendall Schmidt, Brian Bialecki, Sailaja Marella, Cynthia Price, Lisa Cimino, Charles Apgar, Prashant Shah, Bjoern Menze, Jill S Barnholtz-Sloan, Jason Martin, Spyridon Bakas
Although machine learning (ML) has shown promise in numerous domains, there are concerns about generalizability to out-of-sample data.
Potential sources of dataset bias complicate investigation of underdiagnosis by machine learning algorithms
no code implementations • 19 Jan 2022 • Mélanie Bernhardt, Charles Jones, Ben Glocker
The models consistently yield higher FPR on subgroups known to be historically underserved, and the study concludes that the models exhibit and potentially even amplify systematic underdiagnosis.
CrossMoDA 2021 challenge: Benchmark of Cross-Modality Domain Adaptation techniques for Vestibular Schwannoma and Cochlea Segmentation
3 code implementations • 8 Jan 2022 • Reuben Dorent, Aaron Kujawa, Marina Ivory, Spyridon Bakas, Nicola Rieke, Samuel Joutard, Ben Glocker, Jorge Cardoso, Marc Modat, Kayhan Batmanghelich, Arseniy Belkov, Maria Baldeon Calisto, Jae Won Choi, Benoit M. Dawant, Hexin Dong, Sergio Escalera, Yubo Fan, Lasse Hansen, Mattias P. Heinrich, Smriti Joshi, Victoriya Kashtanova, Hyeon Gyu Kim, Satoshi Kondo, Christian N. Kruse, Susana K. Lai-Yuen, Hao Li, Han Liu, Buntheng Ly, Ipek Oguz, Hyungseob Shin, Boris Shirokikh, Zixian Su, Guotai Wang, Jianghao Wu, Yanwu Xu, Kai Yao, Li Zhang, Sebastien Ourselin, Jonathan Shapey, Tom Vercauteren
The aim was to automatically perform unilateral VS and bilateral cochlea segmentation on hrT2 as provided in the testing set (N=137).
A Variational Bayesian Method for Similarity Learning in Non-Rigid Image Registration
1 code implementation • CVPR 2022 • Daniel Grzech, Mohammad Farid Azampour, Ben Glocker, Julia Schnabel, Nassir Navab, Bernhard Kainz, Loïc le Folgoc
We propose a novel variational Bayesian formulation for diffeomorphic non-rigid registration of medical images, which learns in an unsupervised way a data-specific similarity metric.
Matrix Inversion free variational inference in Conditional Student's T Processes
no code implementations • pproximateinference AABI Symposium 2022 • Sebastian Popescu, Ben Glocker, Mark van der Wilk
We propose a new variational lower bound for performing inference in sparse Student's T Processes that does not require computationally intensive operations such as matrix inversions or log determinants of matrices.
Algorithmic encoding of protected characteristics in image-based models for disease detection
1 code implementation • 27 Oct 2021 • Ben Glocker, Charles Jones, Melanie Bernhardt, Stefan Winzeck
We explore test set resampling, transfer learning, multitask learning, and model inspection to assess the relationship between the encoding of protected characteristics and disease detection performance across subgroups.
Uncertainty quantification in non-rigid image registration via stochastic gradient Markov chain Monte Carlo
1 code implementation • 25 Oct 2021 • Daniel Grzech, Mohammad Farid Azampour, Huaqi Qiu, Ben Glocker, Bernhard Kainz, Loïc le Folgoc
We develop a new Bayesian model for non-rigid registration of three-dimensional medical images, with a focus on uncertainty quantification.
Is MC Dropout Bayesian?
no code implementations • 8 Oct 2021 • Loic Le Folgoc, Vasileios Baltatzis, Sujal Desai, Anand Devaraj, Sam Ellis, Octavio E. Martinez Manzanera, Arjun Nair, Huaqi Qiu, Julia Schnabel, Ben Glocker
We question the properties of MC Dropout for approximate inference, as in fact MC Dropout changes the Bayesian model; its predictive posterior assigns $0$ probability to the true model on closed-form benchmarks; the multimodality of its predictive posterior is not a property of the true predictive posterior but a design artefact.
DeepMCAT: Large-Scale Deep Clustering for Medical Image Categorization
no code implementations • 30 Sep 2021 • Turkay Kart, Wenjia Bai, Ben Glocker, Daniel Rueckert
In recent years, the research landscape of machine learning in medical imaging has changed drastically from supervised to semi-, weakly- or unsupervised methods.
Class-Distribution-Aware Calibration for Long-Tailed Visual Recognition
1 code implementation • 11 Sep 2021 • Mobarakol Islam, Lalithkumar Seenivasan, Hongliang Ren, Ben Glocker
In CDA-TS, the scalar temperature value is replaced with the CDA temperature vector encoded with class frequency to compensate for the over-confidence.
Active label cleaning for improved dataset quality under resource constraints
1 code implementation • 1 Sep 2021 • Melanie Bernhardt, Daniel C. Castro, Ryutaro Tanno, Anton Schwaighofer, Kerem C. Tezcan, Miguel Monteiro, Shruthi Bannur, Matthew Lungren, Aditya Nori, Ben Glocker, Javier Alvarez-Valle, Ozan Oktay
Imperfections in data annotation, known as label noise, are detrimental to the training of machine learning models and have an often-overlooked confounding effect on the assessment of model performance.
The Pitfalls of Sample Selection: A Case Study on Lung Nodule Classification
no code implementations • 11 Aug 2021 • Vasileios Baltatzis, Kyriaki-Margarita Bintsi, Loic Le Folgoc, Octavio E. Martinez Manzanera, Sam Ellis, Arjun Nair, Sujal Desai, Ben Glocker, Julia A. Schnabel
Using publicly available data to determine the performance of methodological contributions is important as it facilitates reproducibility and allows scrutiny of the published results.
The Effect of the Loss on Generalization: Empirical Study on Synthetic Lung Nodule Data
no code implementations • 10 Aug 2021 • Vasileios Baltatzis, Loic Le Folgoc, Sam Ellis, Octavio E. Martinez Manzanera, Kyriaki-Margarita Bintsi, Arjun Nair, Sujal Desai, Ben Glocker, Julia A. Schnabel
Convolutional Neural Networks (CNNs) are widely used for image classification in a variety of fields, including medical imaging.
Bayesian analysis of the prevalence bias: learning and predicting from imbalanced data
no code implementations • 31 Jul 2021 • Loic Le Folgoc, Vasileios Baltatzis, Amir Alansary, Sujal Desai, Anand Devaraj, Sam Ellis, Octavio E. Martinez Manzanera, Fahdi Kanavati, Arjun Nair, Julia Schnabel, Ben Glocker
This mismatch is known as sampling bias.
Data synthesis and adversarial networks: A review and meta-analysis in cancer imaging
no code implementations • 20 Jul 2021 • Richard Osuala, Kaisar Kushibar, Lidia Garrucho, Akis Linardos, Zuzanna Szafranowska, Stefan Klein, Ben Glocker, Oliver Diaz, Karim Lekadir
Despite technological and medical advances, the detection, interpretation, and treatment of cancer based on imaging data continue to pose significant challenges.
Transductive image segmentation: Self-training and effect of uncertainty estimation
no code implementations • 19 Jul 2021 • Konstantinos Kamnitsas, Stefan Winzeck, Evgenios N. Kornaropoulos, Daniel Whitehouse, Cameron Englman, Poe Phyu, Norman Pao, David K. Menon, Daniel Rueckert, Tilak Das, Virginia F. J. Newcombe, Ben Glocker
It focuses on the quality of predictions made on the unlabeled data of interest when they are included for optimization during training, rather than improving generalization.
Multiple Instance Learning with Auxiliary Task Weighting for Multiple Myeloma Classification
1 code implementation • 16 Jul 2021 • Talha Qaiser, Stefan Winzeck, Theodore Barfoot, Tara Barwick, Simon J. Doran, Martin F. Kaiser, Linda Wedlake, Nina Tunariu, Dow-Mu Koh, Christina Messiou, Andrea Rockall, Ben Glocker
To aid radiological reading, we propose an auxiliary task-based multiple instance learning approach (ATMIL) for MM classification with the ability to localize sites of disease.
Learning from Partially Overlapping Labels: Image Segmentation under Annotation Shift
no code implementations • 13 Jul 2021 • Gregory Filbrandt, Konstantinos Kamnitsas, David Bernstein, Alexandra Taylor, Ben Glocker
Scarcity of high quality annotated images remains a limiting factor for training accurate image segmentation models.
Detecting Hypo-plastic Left Heart Syndrome in Fetal Ultrasound via Disease-specific Atlas Maps
no code implementations • 6 Jul 2021 • Samuel Budd, Matthew Sinclair, Thomas Day, Athanasios Vlontzos, Jeremy Tan, Tianrui Liu, Jaqueline Matthew, Emily Skelton, John Simpson, Reza Razavi, Ben Glocker, Daniel Rueckert, Emma C. Robinson, Bernhard Kainz
Fetal ultrasound screening during pregnancy plays a vital role in the early detection of fetal malformations which have potential long-term health impacts.
Confidence-based Out-of-Distribution Detection: A Comparative Study and Analysis
1 code implementation • 6 Jul 2021 • Christoph Berger, Magdalini Paschali, Ben Glocker, Konstantinos Kamnitsas
Image classification models deployed in the real world may receive inputs outside the intended data distribution.
Distributional Gaussian Process Layers for Outlier Detection in Image Segmentation
no code implementations • 28 Apr 2021 • Sebastian G. Popescu, David J. Sharp, James H. Cole, Konstantinos Kamnitsas, Ben Glocker
We propose a parameter efficient Bayesian layer for hierarchical convolutional Gaussian Processes that incorporates Gaussian Processes operating in Wasserstein-2 space to reliably propagate uncertainty.
Common Limitations of Image Processing Metrics: A Picture Story
1 code implementation • 12 Apr 2021 • Annika Reinke, Minu D. Tizabi, Carole H. Sudre, Matthias Eisenmann, Tim Rädsch, Michael Baumgartner, Laura Acion, Michela Antonelli, Tal Arbel, Spyridon Bakas, Peter Bankhead, Arriel Benis, Matthew Blaschko, Florian Buettner, M. Jorge Cardoso, Jianxu Chen, Veronika Cheplygina, Evangelia Christodoulou, Beth Cimini, Gary S. Collins, Sandy Engelhardt, Keyvan Farahani, Luciana Ferrer, Adrian Galdran, Bram van Ginneken, Ben Glocker, Patrick Godau, Robert Haase, Fred Hamprecht, Daniel A. Hashimoto, Doreen Heckmann-Nötzel, Peter Hirsch, Michael M. Hoffman, Merel Huisman, Fabian Isensee, Pierre Jannin, Charles E. Kahn, Dagmar Kainmueller, Bernhard Kainz, Alexandros Karargyris, Alan Karthikesalingam, A. Emre Kavur, Hannes Kenngott, Jens Kleesiek, Andreas Kleppe, Sven Kohler, Florian Kofler, Annette Kopp-Schneider, Thijs Kooi, Michal Kozubek, Anna Kreshuk, Tahsin Kurc, Bennett A. Landman, Geert Litjens, Amin Madani, Klaus Maier-Hein, Anne L. Martel, Peter Mattson, Erik Meijering, Bjoern Menze, David Moher, Karel G. M. Moons, Henning Müller, Brennan Nichyporuk, Felix Nickel, M. Alican Noyan, Jens Petersen, Gorkem Polat, Susanne M. Rafelski, Nasir Rajpoot, Mauricio Reyes, Nicola Rieke, Michael Riegler, Hassan Rivaz, Julio Saez-Rodriguez, Clara I. Sánchez, Julien Schroeter, Anindo Saha, M. Alper Selver, Lalith Sharan, Shravya Shetty, Maarten van Smeden, Bram Stieltjes, Ronald M. Summers, Abdel A. Taha, Aleksei Tiulpin, Sotirios A. Tsaftaris, Ben van Calster, Gaël Varoquaux, Manuel Wiesenfarth, Ziv R. Yaniv, Paul Jäger, Lena Maier-Hein
While the importance of automatic image analysis is continuously increasing, recent meta-research revealed major flaws with respect to algorithm validation.
Spatially Varying Label Smoothing: Capturing Uncertainty from Expert Annotations
1 code implementation • 12 Apr 2021 • Mobarakol Islam, Ben Glocker
The task of image segmentation is inherently noisy due to ambiguities regarding the exact location of boundaries between anatomical structures.
Analyzing Overfitting under Class Imbalance in Neural Networks for Image Segmentation
1 code implementation • 20 Feb 2021 • Zeju Li, Konstantinos Kamnitsas, Ben Glocker
In particular, in image segmentation neural networks may overfit to the foreground samples from small structures, which are often heavily under-represented in the training set, leading to poor generalization.
Atlas-ISTN: Joint Segmentation, Registration and Atlas Construction with Image-and-Spatial Transformer Networks
no code implementations • 18 Dec 2020 • Matthew Sinclair, Andreas Schuh, Karl Hahn, Kersten Petersen, Ying Bai, James Batten, Michiel Schaap, Ben Glocker
We propose Atlas-ISTN, a framework that jointly learns segmentation and registration on 2D and 3D image data, and constructs a population-derived atlas in the process.
Decoupled Sparse Gaussian Processes Components]{Decoupled Sparse Gaussian Processes Components : Separating Decision Making from Data Manifold Fitting
no code implementations • pproximateinference AABI Symposium 2021 • Sebastian Popescu, David J. Sharp, James H. Cole, Ben Glocker
We propose a decoupling in Reproducing Kernel Hilbert Space of the parametric and non-parametric components of Sparse Gaussian Processes.
Hierarchical Gaussian Processes with Wasserstein-2 Kernels
no code implementations • 28 Oct 2020 • Sebastian Popescu, David Sharp, James Cole, Ben Glocker
Stacking Gaussian Processes severely diminishes the model's ability to detect outliers, which when combined with non-zero mean functions, further extrapolates low non-parametric variance to low training data density regions.
Cranial Implant Design via Virtual Craniectomy with Shape Priors
no code implementations • 29 Sep 2020 • Franco Matzkin, Virginia Newcombe, Ben Glocker, Enzo Ferrante
Our direct estimation method outperforms the baselines provided by the organizers, while the model with shape priors shows superior performance when dealing with out-of-distribution cases.
Self-supervised Skull Reconstruction in Brain CT Images with Decompressive Craniectomy
1 code implementation • 7 Jul 2020 • Franco Matzkin, Virginia Newcombe, Susan Stevenson, Aneesh Khetani, Tom Newman, Richard Digby, Andrew Stevens, Ben Glocker, Enzo Ferrante
Decompressive craniectomy (DC) is a common surgical procedure consisting of the removal of a portion of the skull that is performed after incidents such as stroke, traumatic brain injury (TBI) or other events that could result in acute subdural hemorrhage and/or increasing intracranial pressure.
Post-DAE: Anatomically Plausible Segmentation via Post-Processing with Denoising Autoencoders
1 code implementation • 24 Jun 2020 • Agostina J. Larrazabal, César Martínez, Ben Glocker, Enzo Ferrante
We introduce Post-DAE, a post-processing method based on denoising autoencoders (DAE) to improve the anatomical plausibility of arbitrary biomedical image segmentation algorithms.
Deep Generative Model-based Quality Control for Cardiac MRI Segmentation
no code implementations • 23 Jun 2020 • Shuo Wang, Giacomo Tarroni, Chen Qin, Yuanhan Mo, Chengliang Dai, Chen Chen, Ben Glocker, Yike Guo, Daniel Rueckert, Wenjia Bai
Our approach provides a real-time and model-agnostic quality control for cardiac MRI segmentation, which has the potential to be integrated into clinical image analysis workflows.
Deep Structural Causal Models for Tractable Counterfactual Inference
3 code implementations • NeurIPS 2020 • Nick Pawlowski, Daniel C. Castro, Ben Glocker
We formulate a general framework for building structural causal models (SCMs) with deep learning components.
Stochastic Segmentation Networks: Modelling Spatially Correlated Aleatoric Uncertainty
1 code implementation • NeurIPS 2020 • Miguel Monteiro, Loïc le Folgoc, Daniel Coelho de Castro, Nick Pawlowski, Bernardo Marques, Konstantinos Kamnitsas, Mark van der Wilk, Ben Glocker
In image segmentation, there is often more than one plausible solution for a given input.
VerSe: A Vertebrae Labelling and Segmentation Benchmark for Multi-detector CT Images
2 code implementations • 24 Jan 2020 • Anjany Sekuboyina, Malek E. Husseini, Amirhossein Bayat, Maximilian Löffler, Hans Liebl, Hongwei Li, Giles Tetteh, Jan Kukačka, Christian Payer, Darko Štern, Martin Urschler, Maodong Chen, Dalong Cheng, Nikolas Lessmann, Yujin Hu, Tianfu Wang, Dong Yang, Daguang Xu, Felix Ambellan, Tamaz Amiranashvili, Moritz Ehlke, Hans Lamecker, Sebastian Lehnert, Marilia Lirio, Nicolás Pérez de Olaguer, Heiko Ramm, Manish Sahu, Alexander Tack, Stefan Zachow, Tao Jiang, Xinjun Ma, Christoph Angerman, Xin Wang, Kevin Brown, Alexandre Kirszenberg, Élodie Puybareau, Di Chen, Yiwei Bai, Brandon H. Rapazzo, Timyoas Yeah, Amber Zhang, Shangliang Xu, Feng Hou, Zhiqiang He, Chan Zeng, Zheng Xiangshang, Xu Liming, Tucker J. Netherton, Raymond P. Mumme, Laurence E. Court, Zixun Huang, Chenhang He, Li-Wen Wang, Sai Ho Ling, Lê Duy Huynh, Nicolas Boutry, Roman Jakubicek, Jiri Chmelik, Supriti Mulay, Mohanasankar Sivaprakasam, Johannes C. Paetzold, Suprosanna Shit, Ivan Ezhov, Benedikt Wiestler, Ben Glocker, Alexander Valentinitsch, Markus Rempfler, Björn H. Menze, Jan S. Kirschke
Two datasets containing a total of 374 multi-detector CT scans from 355 patients were prepared and 4505 vertebrae have individually been annotated at voxel-level by a human-machine hybrid algorithm (https://osf. io/nqjyw/, https://osf. io/t98fz/).
Unpaired Multi-modal Segmentation via Knowledge Distillation
1 code implementation • 6 Jan 2020 • Qi Dou, Quande Liu, Pheng Ann Heng, Ben Glocker
We propose a novel learning scheme for unpaired cross-modality image segmentation, with a highly compact architecture achieving superior segmentation accuracy.
Causality matters in medical imaging
no code implementations • 17 Dec 2019 • Daniel C. Castro, Ian Walker, Ben Glocker
This article discusses how the language of causality can shed new light on the major challenges in machine learning for medical imaging: 1) data scarcity, which is the limited availability of high-quality annotations, and 2) data mismatch, whereby a trained algorithm may fail to generalize in clinical practice.
Universal Adversarial Robustness of Texture and Shape-Biased Models
1 code implementation • 23 Nov 2019 • Kenneth T. Co, Luis Muñoz-González, Leslie Kanthan, Ben Glocker, Emil C. Lupu
Increasing shape-bias in deep neural networks has been shown to improve robustness to common corruptions and noise.
Domain Generalization via Model-Agnostic Learning of Semantic Features
1 code implementation • NeurIPS 2019 • Qi Dou, Daniel C. Castro, Konstantinos Kamnitsas, Ben Glocker
Generalization capability to unseen domains is crucial for machine learning models when deploying to real-world conditions.
 Ranked #66 on
Domain Generalization
on PACS
Ranked #66 on
Domain Generalization
on PACS
Vertebrae Detection and Localization in CT with Two-Stage CNNs and Dense Annotations
1 code implementation • 14 Oct 2019 • James McCouat, Ben Glocker
We propose a new, two-stage approach to the vertebrae centroid detection and localization problem.
Machine Learning with Multi-Site Imaging Data: An Empirical Study on the Impact of Scanner Effects
no code implementations • 10 Oct 2019 • Ben Glocker, Robert Robinson, Daniel C. Castro, Qi Dou, Ender Konukoglu
This is an empirical study to investigate the impact of scanner effects when using machine learning on multi-site neuroimaging data.
Needles in Haystacks: On Classifying Tiny Objects in Large Images
1 code implementation • 16 Aug 2019 • Nick Pawlowski, Suvrat Bhooshan, Nicolas Ballas, Francesco Ciompi, Ben Glocker, Michal Drozdzal
In some important computer vision domains, such as medical or hyperspectral imaging, we care about the classification of tiny objects in large images.
Overfitting of neural nets under class imbalance: Analysis and improvements for segmentation
1 code implementation • 25 Jul 2019 • Zeju Li, Konstantinos Kamnitsas, Ben Glocker
Overfitting in deep learning has been the focus of a number of recent works, yet its exact impact on the behavior of neural networks is not well understood.
Is Texture Predictive for Age and Sex in Brain MRI?
1 code implementation • 25 Jul 2019 • Nick Pawlowski, Ben Glocker
Deep learning builds the foundation for many medical image analysis tasks where neuralnetworks are often designed to have a large receptive field to incorporate long spatialdependencies.
Image-and-Spatial Transformer Networks for Structure-Guided Image Registration
1 code implementation • 22 Jul 2019 • Matthew C. H. Lee, Ozan Oktay, Andreas Schuh, Michiel Schaap, Ben Glocker
The goal is to learn a complex function that maps the appearance of input image pairs to parameters of a spatial transformation in order to align corresponding anatomical structures.
Improving RetinaNet for CT Lesion Detection with Dense Masks from Weak RECIST Labels
3 code implementations • 5 Jun 2019 • Martin Zlocha, Qi Dou, Ben Glocker
We propose a highly accurate and efficient one-stage lesion detector, by re-designing a RetinaNet to meet the particular challenges in medical imaging.
 Ranked #9 on
Medical Object Detection
on DeepLesion
Ranked #9 on
Medical Object Detection
on DeepLesion
Medical Imaging with Deep Learning: MIDL 2019 -- Extended Abstract Track
no code implementations • 21 May 2019 • M. Jorge Cardoso, Aasa Feragen, Ben Glocker, Ender Konukoglu, Ipek Oguz, Gozde Unal, Tom Vercauteren
This compendium gathers all the accepted extended abstracts from the Second International Conference on Medical Imaging with Deep Learning (MIDL 2019), held in London, UK, 8-10 July 2019.
Quantitative Error Prediction of Medical Image Registration using Regression Forests
1 code implementation • 18 May 2019 • Hessam Sokooti, Gorkem Saygili, Ben Glocker, Boudewijn P. F. Lelieveldt, Marius Staring
This paper proposes a new automatic method to predict the registration error in a quantitative manner, and is applied to chest CT scans.
Graph Convolutional Gaussian Processes
no code implementations • 14 May 2019 • Ian Walker, Ben Glocker
We propose a novel Bayesian nonparametric method to learn translation-invariant relationships on non-Euclidean domains.
FastReg: Fast Non-Rigid Registration via Accelerated Optimisation on the Manifold of Diffeomorphisms
no code implementations • 5 Mar 2019 • Daniel Grzech, Loïc le Folgoc, Mattias P. Heinrich, Bishesh Khanal, Jakub Moll, Julia A. Schnabel, Ben Glocker, Bernhard Kainz
We present an implementation of a new approach to diffeomorphic non-rigid registration of medical images.
Automated Quality Control in Image Segmentation: Application to the UK Biobank Cardiac MR Imaging Study
no code implementations • 27 Jan 2019 • Robert Robinson, Vanya V. Valindria, Wenjia Bai, Ozan Oktay, Bernhard Kainz, Hideaki Suzuki, Mihir M. Sanghvi, Nay Aung, Jos$é$ Miguel Paiva, Filip Zemrak, Kenneth Fung, Elena Lukaschuk, Aaron M. Lee, Valentina Carapella, Young Jin Kim, Stefan K. Piechnik, Stefan Neubauer, Steffen E. Petersen, Chris Page, Paul M. Matthews, Daniel Rueckert, Ben Glocker
Methods: To overcome this challenge, we explore an approach for predicting segmentation quality based on Reverse Classification Accuracy, which enables us to discriminate between successful and failed segmentations on a per-cases basis.
PnP-AdaNet: Plug-and-Play Adversarial Domain Adaptation Network with a Benchmark at Cross-modality Cardiac Segmentation
2 code implementations • 19 Dec 2018 • Qi Dou, Cheng Ouyang, Cheng Chen, Hao Chen, Ben Glocker, Xiahai Zhuang, Pheng-Ann Heng
In this paper, we propose the PnPAdaNet (plug-and-play adversarial domain adaptation network) for adapting segmentation networks between different modalities of medical images, e. g., MRI and CT. We propose to tackle the significant domain shift by aligning the feature spaces of source and target domains in an unsupervised manner.
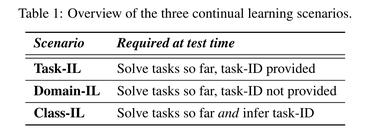
Towards continual learning in medical imaging
no code implementations • 6 Nov 2018 • Chaitanya Baweja, Ben Glocker, Konstantinos Kamnitsas
This work investigates continual learning of two segmentation tasks in brain MRI with neural networks.
Identifying the Best Machine Learning Algorithms for Brain Tumor Segmentation, Progression Assessment, and Overall Survival Prediction in the BRATS Challenge
1 code implementation • 5 Nov 2018 • Spyridon Bakas, Mauricio Reyes, Andras Jakab, Stefan Bauer, Markus Rempfler, Alessandro Crimi, Russell Takeshi Shinohara, Christoph Berger, Sung Min Ha, Martin Rozycki, Marcel Prastawa, Esther Alberts, Jana Lipkova, John Freymann, Justin Kirby, Michel Bilello, Hassan Fathallah-Shaykh, Roland Wiest, Jan Kirschke, Benedikt Wiestler, Rivka Colen, Aikaterini Kotrotsou, Pamela Lamontagne, Daniel Marcus, Mikhail Milchenko, Arash Nazeri, Marc-Andre Weber, Abhishek Mahajan, Ujjwal Baid, Elizabeth Gerstner, Dongjin Kwon, Gagan Acharya, Manu Agarwal, Mahbubul Alam, Alberto Albiol, Antonio Albiol, Francisco J. Albiol, Varghese Alex, Nigel Allinson, Pedro H. A. Amorim, Abhijit Amrutkar, Ganesh Anand, Simon Andermatt, Tal Arbel, Pablo Arbelaez, Aaron Avery, Muneeza Azmat, Pranjal B., W Bai, Subhashis Banerjee, Bill Barth, Thomas Batchelder, Kayhan Batmanghelich, Enzo Battistella, Andrew Beers, Mikhail Belyaev, Martin Bendszus, Eze Benson, Jose Bernal, Halandur Nagaraja Bharath, George Biros, Sotirios Bisdas, James Brown, Mariano Cabezas, Shilei Cao, Jorge M. Cardoso, Eric N Carver, Adrià Casamitjana, Laura Silvana Castillo, Marcel Catà, Philippe Cattin, Albert Cerigues, Vinicius S. Chagas, Siddhartha Chandra, Yi-Ju Chang, Shiyu Chang, Ken Chang, Joseph Chazalon, Shengcong Chen, Wei Chen, Jefferson W. Chen, Zhaolin Chen, Kun Cheng, Ahana Roy Choudhury, Roger Chylla, Albert Clérigues, Steven Colleman, Ramiro German Rodriguez Colmeiro, Marc Combalia, Anthony Costa, Xiaomeng Cui, Zhenzhen Dai, Lutao Dai, Laura Alexandra Daza, Eric Deutsch, Changxing Ding, Chao Dong, Shidu Dong, Wojciech Dudzik, Zach Eaton-Rosen, Gary Egan, Guilherme Escudero, Théo Estienne, Richard Everson, Jonathan Fabrizio, Yong Fan, Longwei Fang, Xue Feng, Enzo Ferrante, Lucas Fidon, Martin Fischer, Andrew P. French, Naomi Fridman, Huan Fu, David Fuentes, Yaozong Gao, Evan Gates, David Gering, Amir Gholami, Willi Gierke, Ben Glocker, Mingming Gong, Sandra González-Villá, T. Grosges, Yuanfang Guan, Sheng Guo, Sudeep Gupta, Woo-Sup Han, Il Song Han, Konstantin Harmuth, Huiguang He, Aura Hernández-Sabaté, Evelyn Herrmann, Naveen Himthani, Winston Hsu, Cheyu Hsu, Xiaojun Hu, Xiaobin Hu, Yan Hu, Yifan Hu, Rui Hua, Teng-Yi Huang, Weilin Huang, Sabine Van Huffel, Quan Huo, Vivek HV, Khan M. Iftekharuddin, Fabian Isensee, Mobarakol Islam, Aaron S. Jackson, Sachin R. Jambawalikar, Andrew Jesson, Weijian Jian, Peter Jin, V Jeya Maria Jose, Alain Jungo, B Kainz, Konstantinos Kamnitsas, Po-Yu Kao, Ayush Karnawat, Thomas Kellermeier, Adel Kermi, Kurt Keutzer, Mohamed Tarek Khadir, Mahendra Khened, Philipp Kickingereder, Geena Kim, Nik King, Haley Knapp, Urspeter Knecht, Lisa Kohli, Deren Kong, Xiangmao Kong, Simon Koppers, Avinash Kori, Ganapathy Krishnamurthi, Egor Krivov, Piyush Kumar, Kaisar Kushibar, Dmitrii Lachinov, Tryphon Lambrou, Joon Lee, Chengen Lee, Yuehchou Lee, M Lee, Szidonia Lefkovits, Laszlo Lefkovits, James Levitt, Tengfei Li, Hongwei Li, Hongyang Li, Xiaochuan Li, Yuexiang Li, Heng Li, Zhenye Li, Xiaoyu Li, Zeju Li, Xiaogang Li, Wenqi Li, Zheng-Shen Lin, Fengming Lin, Pietro Lio, Chang Liu, Boqiang Liu, Xiang Liu, Mingyuan Liu, Ju Liu, Luyan Liu, Xavier Llado, Marc Moreno Lopez, Pablo Ribalta Lorenzo, Zhentai Lu, Lin Luo, Zhigang Luo, Jun Ma, Kai Ma, Thomas Mackie, Anant Madabushi, Issam Mahmoudi, Klaus H. Maier-Hein, Pradipta Maji, CP Mammen, Andreas Mang, B. S. Manjunath, Michal Marcinkiewicz, S McDonagh, Stephen McKenna, Richard McKinley, Miriam Mehl, Sachin Mehta, Raghav Mehta, Raphael Meier, Christoph Meinel, Dorit Merhof, Craig Meyer, Robert Miller, Sushmita Mitra, Aliasgar Moiyadi, David Molina-Garcia, Miguel A. B. Monteiro, Grzegorz Mrukwa, Andriy Myronenko, Jakub Nalepa, Thuyen Ngo, Dong Nie, Holly Ning, Chen Niu, Nicholas K Nuechterlein, Eric Oermann, Arlindo Oliveira, Diego D. C. Oliveira, Arnau Oliver, Alexander F. I. Osman, Yu-Nian Ou, Sebastien Ourselin, Nikos Paragios, Moo Sung Park, Brad Paschke, J. Gregory Pauloski, Kamlesh Pawar, Nick Pawlowski, Linmin Pei, Suting Peng, Silvio M. Pereira, Julian Perez-Beteta, Victor M. Perez-Garcia, Simon Pezold, Bao Pham, Ashish Phophalia, Gemma Piella, G. N. Pillai, Marie Piraud, Maxim Pisov, Anmol Popli, Michael P. Pound, Reza Pourreza, Prateek Prasanna, Vesna Prkovska, Tony P. Pridmore, Santi Puch, Élodie Puybareau, Buyue Qian, Xu Qiao, Martin Rajchl, Swapnil Rane, Michael Rebsamen, Hongliang Ren, Xuhua Ren, Karthik Revanuru, Mina Rezaei, Oliver Rippel, Luis Carlos Rivera, Charlotte Robert, Bruce Rosen, Daniel Rueckert, Mohammed Safwan, Mostafa Salem, Joaquim Salvi, Irina Sanchez, Irina Sánchez, Heitor M. Santos, Emmett Sartor, Dawid Schellingerhout, Klaudius Scheufele, Matthew R. Scott, Artur A. Scussel, Sara Sedlar, Juan Pablo Serrano-Rubio, N. Jon Shah, Nameetha Shah, Mazhar Shaikh, B. Uma Shankar, Zeina Shboul, Haipeng Shen, Dinggang Shen, Linlin Shen, Haocheng Shen, Varun Shenoy, Feng Shi, Hyung Eun Shin, Hai Shu, Diana Sima, M Sinclair, Orjan Smedby, James M. Snyder, Mohammadreza Soltaninejad, Guidong Song, Mehul Soni, Jean Stawiaski, Shashank Subramanian, Li Sun, Roger Sun, Jiawei Sun, Kay Sun, Yu Sun, Guoxia Sun, Shuang Sun, Yannick R Suter, Laszlo Szilagyi, Sanjay Talbar, DaCheng Tao, Zhongzhao Teng, Siddhesh Thakur, Meenakshi H Thakur, Sameer Tharakan, Pallavi Tiwari, Guillaume Tochon, Tuan Tran, Yuhsiang M. Tsai, Kuan-Lun Tseng, Tran Anh Tuan, Vadim Turlapov, Nicholas Tustison, Maria Vakalopoulou, Sergi Valverde, Rami Vanguri, Evgeny Vasiliev, Jonathan Ventura, Luis Vera, Tom Vercauteren, C. A. Verrastro, Lasitha Vidyaratne, Veronica Vilaplana, Ajeet Vivekanandan, Qian Wang, Chiatse J. Wang, Wei-Chung Wang, Duo Wang, Ruixuan Wang, Yuanyuan Wang, Chunliang Wang, Guotai Wang, Ning Wen, Xin Wen, Leon Weninger, Wolfgang Wick, Shaocheng Wu, Qiang Wu, Yihong Wu, Yong Xia, Yanwu Xu, Xiaowen Xu, Peiyuan Xu, Tsai-Ling Yang, Xiaoping Yang, Hao-Yu Yang, Junlin Yang, Haojin Yang, Guang Yang, Hongdou Yao, Xujiong Ye, Changchang Yin, Brett Young-Moxon, Jinhua Yu, Xiangyu Yue, Songtao Zhang, Angela Zhang, Kun Zhang, Xue-jie Zhang, Lichi Zhang, Xiaoyue Zhang, Yazhuo Zhang, Lei Zhang, Jian-Guo Zhang, Xiang Zhang, Tianhao Zhang, Sicheng Zhao, Yu Zhao, Xiaomei Zhao, Liang Zhao, Yefeng Zheng, Liming Zhong, Chenhong Zhou, Xiaobing Zhou, Fan Zhou, Hongtu Zhu, Jin Zhu, Ying Zhuge, Weiwei Zong, Jayashree Kalpathy-Cramer, Keyvan Farahani, Christos Davatzikos, Koen van Leemput, Bjoern Menze
This study assesses the state-of-the-art machine learning (ML) methods used for brain tumor image analysis in mpMRI scans, during the last seven instances of the International Brain Tumor Segmentation (BraTS) challenge, i. e., 2012-2018.
Morpho-MNIST: Quantitative Assessment and Diagnostics for Representation Learning
1 code implementation • ICLR 2019 • Daniel C. Castro, Jeremy Tan, Bernhard Kainz, Ender Konukoglu, Ben Glocker
Revealing latent structure in data is an active field of research, having introduced exciting technologies such as variational autoencoders and adversarial networks, and is essential to push machine learning towards unsupervised knowledge discovery.
Attention Gated Networks: Learning to Leverage Salient Regions in Medical Images
2 code implementations • 22 Aug 2018 • Jo Schlemper, Ozan Oktay, Michiel Schaap, Mattias Heinrich, Bernhard Kainz, Ben Glocker, Daniel Rueckert
AGs can be easily integrated into standard CNN models such as VGG or U-Net architectures with minimal computational overhead while increasing the model sensitivity and prediction accuracy.
Small Organ Segmentation in Whole-body MRI using a Two-stage FCN and Weighting Schemes
no code implementations • 30 Jul 2018 • Vanya V. Valindria, Ioannis Lavdas, Juan Cerrolaza, Eric O. Aboagye, Andrea G. Rockall, Daniel Rueckert, Ben Glocker
Our experiments show that the proposed approach can boost the segmentation accuracy of multiple small organs in whole-body MRI scans.
High-Resolution Mammogram Synthesis using Progressive Generative Adversarial Networks
no code implementations • 9 Jul 2018 • Dimitrios Korkinof, Tobias Rijken, Michael O'Neill, Joseph Yearsley, Hugh Harvey, Ben Glocker
The ability to generate synthetic medical images is useful for data augmentation, domain transfer, and out-of-distribution detection.
Real-time Prediction of Segmentation Quality
no code implementations • 16 Jun 2018 • Robert Robinson, Ozan Oktay, Wenjia Bai, Vanya Valindria, Mihir Sanghvi, Nay Aung, José Paiva, Filip Zemrak, Kenneth Fung, Elena Lukaschuk, Aaron Lee, Valentina Carapella, Young Jin Kim, Bernhard Kainz, Stefan Piechnik, Stefan Neubauer, Steffen Petersen, Chris Page, Daniel Rueckert, Ben Glocker
Recent advances in deep learning based image segmentation methods have enabled real-time performance with human-level accuracy.
Deep Generative Models in the Real-World: An Open Challenge from Medical Imaging
no code implementations • 14 Jun 2018 • Xiaoran Chen, Nick Pawlowski, Martin Rajchl, Ben Glocker, Ender Konukoglu
In this paper, we explore the feasibility of using state-of-the-art auto-encoder-based deep generative models, such as variational and adversarial auto-encoders, for one such task: abnormality detection in medical imaging.
NeuroNet: Fast and Robust Reproduction of Multiple Brain Image Segmentation Pipelines
1 code implementation • 11 Jun 2018 • Martin Rajchl, Nick Pawlowski, Daniel Rueckert, Paul M. Matthews, Ben Glocker
NeuroNet is a deep convolutional neural network mimicking multiple popular and state-of-the-art brain segmentation tools including FSL, SPM, and MALPEM.
Automatic View Planning with Multi-scale Deep Reinforcement Learning Agents
no code implementations • 8 Jun 2018 • Amir Alansary, Loic Le Folgoc, Ghislain Vaillant, Ozan Oktay, Yuanwei Li, Wenjia Bai, Jonathan Passerat-Palmbach, Ricardo Guerrero, Konstantinos Kamnitsas, Benjamin Hou, Steven McDonagh, Ben Glocker, Bernhard Kainz, Daniel Rueckert
Navigating through target anatomy to find the required view plane is tedious and operator-dependent.
Semi-Supervised Learning via Compact Latent Space Clustering
no code implementations • ICML 2018 • Konstantinos Kamnitsas, Daniel C. Castro, Loic Le Folgoc, Ian Walker, Ryutaro Tanno, Daniel Rueckert, Ben Glocker, Antonio Criminisi, Aditya Nori
We present a novel cost function for semi-supervised learning of neural networks that encourages compact clustering of the latent space to facilitate separation.
Nonparametric Density Flows for MRI Intensity Normalisation
1 code implementation • 7 Jun 2018 • Daniel C. Castro, Ben Glocker
With the adoption of powerful machine learning methods in medical image analysis, it is becoming increasingly desirable to aggregate data that is acquired across multiple sites.
Graph Saliency Maps through Spectral Convolutional Networks: Application to Sex Classification with Brain Connectivity
no code implementations • 5 Jun 2018 • Salim Arslan, Sofia Ira Ktena, Ben Glocker, Daniel Rueckert
Graph convolutional networks (GCNs) allow to apply traditional convolution operations in non-Euclidean domains, where data are commonly modelled as irregular graphs.
Disease Prediction using Graph Convolutional Networks: Application to Autism Spectrum Disorder and Alzheimer's Disease
1 code implementation • 5 Jun 2018 • Sarah Parisot, Sofia Ira Ktena, Enzo Ferrante, Matthew Lee, Ricardo Guerrero, Ben Glocker, Daniel Rueckert
Graphs are widely used as a natural framework that captures interactions between individual elements represented as nodes in a graph.
Domain Adaptation for MRI Organ Segmentation using Reverse Classification Accuracy
1 code implementation • 1 Jun 2018 • Vanya V. Valindria, Ioannis Lavdas, Wenjia Bai, Konstantinos Kamnitsas, Eric O. Aboagye, Andrea G. Rockall, Daniel Rueckert, Ben Glocker
The variations in multi-center data in medical imaging studies have brought the necessity of domain adaptation.
Computing CNN Loss and Gradients for Pose Estimation with Riemannian Geometry
1 code implementation • 2 May 2018 • Benjamin Hou, Nina Miolane, Bishesh Khanal, Matthew C. H. Lee, Amir Alansary, Steven McDonagh, Jo V. Hajnal, Daniel Rueckert, Ben Glocker, Bernhard Kainz
In this paper, we propose a general Riemannian formulation of the pose estimation problem.
Attention-Gated Networks for Improving Ultrasound Scan Plane Detection
6 code implementations • 15 Apr 2018 • Jo Schlemper, Ozan Oktay, Liang Chen, Jacqueline Matthew, Caroline Knight, Bernhard Kainz, Ben Glocker, Daniel Rueckert
We show that, when the base network has a high capacity, the incorporated attention mechanism can provide efficient object localisation while improving the overall performance.
Attention U-Net: Learning Where to Look for the Pancreas
36 code implementations • 11 Apr 2018 • Ozan Oktay, Jo Schlemper, Loic Le Folgoc, Matthew Lee, Mattias Heinrich, Kazunari Misawa, Kensaku Mori, Steven McDonagh, Nils Y. Hammerla, Bernhard Kainz, Ben Glocker, Daniel Rueckert
We propose a novel attention gate (AG) model for medical imaging that automatically learns to focus on target structures of varying shapes and sizes.
 Ranked #1 on
Pancreas Segmentation
on CT-150
Ranked #1 on
Pancreas Segmentation
on CT-150
Learning-Based Quality Control for Cardiac MR Images
no code implementations • 25 Mar 2018 • Giacomo Tarroni, Ozan Oktay, Wenjia Bai, Andreas Schuh, Hideaki Suzuki, Jonathan Passerat-Palmbach, Antonio de Marvao, Declan P. O'Regan, Stuart Cook, Ben Glocker, Paul M. Matthews, Daniel Rueckert
The results show the capability of the proposed pipeline to correctly detect incomplete or corrupted scans (e. g. on UK Biobank, sensitivity and specificity respectively 88% and 99% for heart coverage estimation, 85% and 95% for motion detection), allowing their exclusion from the analysed dataset or the triggering of a new acquisition.
DLTK: State of the Art Reference Implementations for Deep Learning on Medical Images
1 code implementation • 18 Nov 2017 • Nick Pawlowski, Sofia Ira Ktena, Matthew C. H. Lee, Bernhard Kainz, Daniel Rueckert, Ben Glocker, Martin Rajchl
We present DLTK, a toolkit providing baseline implementations for efficient experimentation with deep learning methods on biomedical images.
Ensembles of Multiple Models and Architectures for Robust Brain Tumour Segmentation
no code implementations • 4 Nov 2017 • Konstantinos Kamnitsas, Wenjia Bai, Enzo Ferrante, Steven McDonagh, Matthew Sinclair, Nick Pawlowski, Martin Rajchl, Matthew Lee, Bernhard Kainz, Daniel Rueckert, Ben Glocker
Deep learning approaches such as convolutional neural nets have consistently outperformed previous methods on challenging tasks such as dense, semantic segmentation.
Implicit Weight Uncertainty in Neural Networks
2 code implementations • 3 Nov 2017 • Nick Pawlowski, Andrew Brock, Matthew C. H. Lee, Martin Rajchl, Ben Glocker
Modern neural networks tend to be overconfident on unseen, noisy or incorrectly labelled data and do not produce meaningful uncertainty measures.
Automated cardiovascular magnetic resonance image analysis with fully convolutional networks
1 code implementation • 25 Oct 2017 • Wenjia Bai, Matthew Sinclair, Giacomo Tarroni, Ozan Oktay, Martin Rajchl, Ghislain Vaillant, Aaron M. Lee, Nay Aung, Elena Lukaschuk, Mihir M. Sanghvi, Filip Zemrak, Kenneth Fung, Jose Miguel Paiva, Valentina Carapella, Young Jin Kim, Hideaki Suzuki, Bernhard Kainz, Paul M. Matthews, Steffen E. Petersen, Stefan K. Piechnik, Stefan Neubauer, Ben Glocker, Daniel Rueckert
By combining FCN with a large-scale annotated dataset, the proposed automated method achieves a high performance on par with human experts in segmenting the LV and RV on short-axis CMR images and the left atrium (LA) and right atrium (RA) on long-axis CMR images.
3D Reconstruction in Canonical Co-ordinate Space from Arbitrarily Oriented 2D Images
no code implementations • 19 Sep 2017 • Benjamin Hou, Bishesh Khanal, Amir Alansary, Steven McDonagh, Alice Davidson, Mary Rutherford, Jo V. Hajnal, Daniel Rueckert, Ben Glocker, Bernhard Kainz
We extensively evaluate the effectiveness of our approach quantitatively on simulated Magnetic Resonance Imaging (MRI), fetal brain imagery with synthetic motion and further demonstrate qualitative results on real fetal MRI data where our method is integrated into a full reconstruction and motion compensation pipeline.
Anatomically Constrained Neural Networks (ACNN): Application to Cardiac Image Enhancement and Segmentation
1 code implementation • 22 May 2017 • Ozan Oktay, Enzo Ferrante, Konstantinos Kamnitsas, Mattias Heinrich, Wenjia Bai, Jose Caballero, Stuart Cook, Antonio de Marvao, Timothy Dawes, Declan O'Regan, Bernhard Kainz, Ben Glocker, Daniel Rueckert
However, in most recent and promising techniques such as CNN based segmentation it is not obvious how to incorporate such prior knowledge.
Efficient variational Bayesian neural network ensembles for outlier detection
1 code implementation • 20 Mar 2017 • Nick Pawlowski, Miguel Jaques, Ben Glocker
In this work we perform outlier detection using ensembles of neural networks obtained by variational approximation of the posterior in a Bayesian neural network setting.
Spectral Graph Convolutions for Population-based Disease Prediction
1 code implementation • 8 Mar 2017 • Sarah Parisot, Sofia Ira Ktena, Enzo Ferrante, Matthew Lee, Ricardo Guerrerro Moreno, Ben Glocker, Daniel Rueckert
We demonstrate the potential of the method on the challenging ADNI and ABIDE databases, as a proof of concept of the benefit from integrating contextual information in classification tasks.
Distance Metric Learning using Graph Convolutional Networks: Application to Functional Brain Networks
3 code implementations • 7 Mar 2017 • Sofia Ira Ktena, Sarah Parisot, Enzo Ferrante, Martin Rajchl, Matthew Lee, Ben Glocker, Daniel Rueckert
Evaluating similarity between graphs is of major importance in several computer vision and pattern recognition problems, where graph representations are often used to model objects or interactions between elements.
Predicting Slice-to-Volume Transformation in Presence of Arbitrary Subject Motion
1 code implementation • 28 Feb 2017 • Benjamin Hou, Amir Alansary, Steven McDonagh, Alice Davidson, Mary Rutherford, Jo V. Hajnal, Daniel Rueckert, Ben Glocker, Bernhard Kainz
Our approach is attractive in challenging imaging scenarios, where significant subject motion complicates reconstruction performance of 3D volumes from 2D slice data.
Reverse Classification Accuracy: Predicting Segmentation Performance in the Absence of Ground Truth
no code implementations • 11 Feb 2017 • Vanya V. Valindria, Ioannis Lavdas, Wenjia Bai, Konstantinos Kamnitsas, Eric O. Aboagye, Andrea G. Rockall, Daniel Rueckert, Ben Glocker
In RCA we take the predicted segmentation from a new image to train a reverse classifier which is evaluated on a set of reference images with available ground truth.
Reconstructing Subject-Specific Effect Maps
1 code implementation • 10 Jan 2017 • Ender Konukoglu, Ben Glocker
Analyses on the ADNI dataset show that RSM can also improve correlation between subject-specific detections in cortical thickness data and non-imaging markers of Alzheimer's Disease (AD), such as the Mini Mental State Examination Score and Cerebrospinal Fluid amyloid-$\beta$ levels.
Unsupervised domain adaptation in brain lesion segmentation with adversarial networks
1 code implementation • 28 Dec 2016 • Konstantinos Kamnitsas, Christian Baumgartner, Christian Ledig, Virginia F. J. Newcombe, Joanna P. Simpson, Andrew D. Kane, David K. Menon, Aditya Nori, Antonio Criminisi, Daniel Rueckert, Ben Glocker
In this work we investigate unsupervised domain adaptation using adversarial neural networks to train a segmentation method which is more invariant to differences in the input data, and which does not require any annotations on the test domain.
Efficient Multi-Scale 3D CNN with Fully Connected CRF for Accurate Brain Lesion Segmentation
2 code implementations • 18 Mar 2016 • Konstantinos Kamnitsas, Christian Ledig, Virginia F. J. Newcombe, Joanna P. Simpson, Andrew D. Kane, David K. Menon, Daniel Rueckert, Ben Glocker
We propose a dual pathway, 11-layers deep, three-dimensional Convolutional Neural Network for the challenging task of brain lesion segmentation.
 Ranked #1 on
Lesion Segmentation
on ISLES-2015
Ranked #1 on
Lesion Segmentation
on ISLES-2015
 3D Medical Imaging Segmentation
3D Medical Imaging Segmentation
 Brain Lesion Segmentation From Mri
+3
Brain Lesion Segmentation From Mri
+3
Multi-Output Learning for Camera Relocalization
no code implementations • CVPR 2014 • Abner Guzman-Rivera, Pushmeet Kohli, Ben Glocker, Jamie Shotton, Toby Sharp, Andrew Fitzgibbon, Shahram Izadi
We formulate this problem as inversion of the generative rendering procedure, i. e., we want to find the camera pose corresponding to a rendering of the 3D scene model that is most similar with the observed input.
Scene Coordinate Regression Forests for Camera Relocalization in RGB-D Images
no code implementations • CVPR 2013 • Jamie Shotton, Ben Glocker, Christopher Zach, Shahram Izadi, Antonio Criminisi, Andrew Fitzgibbon
We address the problem of inferring the pose of an RGB-D camera relative to a known 3D scene, given only a single acquired image.